AcceleratedLifeModel
Description
An AcceleratedLifeModel object contains the results of fitting an
accelerated life model to stressor and failure time data.
Use the properties of an AcceleratedLifeModel object to investigate a
fitted accelerated life model. The object properties include the stressor and failure time
data, and information about coefficient estimates, statistics, and censoring. Use the object
functions to predict mean failure times at stressor levels, compute confidence intervals for
coefficient estimates, and visualize the accelerated life model. For more information, see
What Is Accelerated Life Analysis?
Creation
Create an AcceleratedLifeModel object using fitacclife.
Properties
This property is read-only.
Stressor levels, represented as a table with n variables, where
n is the number of elements in StressorNames. Each column
(variable) in StressorLevels contains a stressor in the accelerated
life model, and each row contains a stressor level.
This property is read-only.
Failure times, represented as a numeric column vector or a two-column numeric
matrix. Each element in FailureTimes corresponds to the row with
the same index in StressorLevels. When
FailureTimes is a two-column matrix, the first column contains
the lower bounds for the interval-censored failure times, and the second column contains
the upper bounds.
Data Types: double
This property is read-only.
Indicator of censored failure times, represented as a vector consisting of
0, –1, and 1, which indicate
fully observed, left-censored, and right-censored failure times, respectively. Each
element of Censoring indicates the censorship status of the
corresponding failure time in FailureTimes. The default is a vector
of 0s, indicating all failure times are fully observed.
Data Types: logical | single | double
This property is read-only.
Life distribution, represented as a string scalar. The following table shows the supported distributions.
| Distribution Name | Distribution Object |
|---|---|
"exponential" | ExponentialDistribution |
"logistic" | LogisticDistribution |
"loglogistic" | LoglogisticDistribution |
"lognormal" | LognormalDistribution |
"normal" | NormalDistribution |
"weibull" (default) | WeibullDistribution |
For more information about life distributions, see What Is Accelerated Life Analysis?
This property is read-only.
Life stress model, represented as a string scalar or a function handle
(@stressName). The following table shows the supported life stress
models, where X1 is the value of the first
stressor, X2 is the value of the second
stressor (if applicable), and bx are the model
coefficients.
| Value | Life Stress Model | Stressor Data Conditions |
|---|---|---|
"linear" | b0 + b1*X1 +
b2*X2 | None |
"log" | b0 + b1*log(X1)
+
b2*log(X2) | X1 and X2
must be positive. |
"exponential" | b0*exp(b1*X1 +
b2*X2) | None |
"arrhenius" (default) | b0*exp(b1/X1 +
b2/X2) | X1 and X2
must be nonzero. |
"power" | b0*X1b1*X2b2 | X1 and X2
must be nonnegative. |
"reciprocal" | b0 + b1/X1 +
b2/X2 | X1 and X2
must be nonzero. |
"sqrt" | b0 +
b1*sqrt(X1)
+
b2*sqrt(X2) | X1 and X2
must be nonnegative. |
"eyring" | (1/X1)*e–(b0 –
b1/X1) | X1 and X2
must be nonzero. |
"temp-nonthermal" | b0*e-b1/X1*X2b2 | X1 must be nonzero, and
X2 must be nonnegative. |
For more information about life stress models, see What Is Accelerated Life Analysis?
Data Types: string | function handle
This property is read-only.
Coefficient values, represented as a table that contains one row for each coefficient and these columns:
Source— Source of the coefficient value, where"StressModel"indicates a life stress model coefficient, and"Distribution"indicates a life distribution coefficientEstimate— Estimated coefficient valueSE— Standard error of the estimate
Use the coefci
function to find the confidence intervals of the coefficient estimates.
To obtain any of the columns as a vector, index into the property using dot
notation. For example, obtain the estimated coefficient vector in the model
mdl:
beta = mdl.Coefficients.Estimate
This property is read-only.
Covariance matrix for the coefficient estimates, represented as a square matrix with the number of rows equal to the number of coefficients.
Data Types: double
This property is read-only.
Coefficient names, represented as a cell array of character vectors, each containing the name of the corresponding term.
Data Types: cell
This property is read-only.
Loglikelihood of the fitted model, represented as a numeric scalar.
You can use loglikelihood values to compare different models and assess the significance of the effects of terms in the model.
Data Types: double
This property is read-only.
Criterion for model comparison, represented as a structure with these fields:
AIC— Akaike information criterion.AIC = –2*logL + 2*m, wherelogLis the loglikelihood andmis the number of estimated parameters.AICc— Akaike information criterion corrected for the sample size.AICc = AIC + (2*m*(m + 1))/(n – m – 1), wherenis the number of observations.BIC— Bayesian information criterion.BIC = –2*logL + m*log(n).CAIC— Consistent Akaike information criterion.CAIC = –2*logL + m*(log(n) + 1).
Information criteria are model selection tools that you can use to compare multiple models fit to the same data. These criteria are likelihood-based measures of model fit that include a penalty for complexity (specifically, the number of parameters). Different information criteria are distinguished by the form of the penalty.
When you compare multiple models, the model with the lowest information criterion value is the best-fitting model. The best-fitting model can vary depending on the criterion used for model comparison.
To obtain any of the criterion values as a scalar, index into the property using dot
notation. For example, obtain the AIC value aic in the model
mdl:
aic = mdl.ModelCriterion.AIC
Data Types: struct
This property is read-only.
Baseline stressor level, represented as a numeric vector with the same number of
elements as StressorNames.
Data Types: double
This property is read-only.
Stressor names, represented as a string array with n elements,
where n is the number of stressors in the model. Each element of
StressorNames contains the name of the corresponding stressor in
StressorLevels.
Data Types: string
Object Functions
accelfactor | Acceleration factors of accelerated life model |
coefci | Confidence intervals for accelerated life model coefficients |
distfcn | Distribution functions of accelerated life model |
distplot | Plot distribution functions of accelerated life model |
icdf | Inverse cumulative distribution function of accelerated life model |
meanfailplot | Plot failure times of accelerated life model |
meanfailtime | Mean failure times and life distribution coefficients of accelerated life model |
probplot | Plot failure probabilities of accelerated life model |
Examples
Perform an accelerated life analysis on a machine part that is subject to different stress levels.
Load and Visualize Data
Load the machineFailure data set, which contains 100 simulated failure time measurements of the machine part at five evenly spaced temperatures (T) between 300 K and 400 K. Approximately 5% of the failure time measurements are right-censored. These measurements are indicated in the censored variable of the table.
load machineFailureCreate a histogram of the failure times at the lowest stressor level in the data set (T = 300).
histogram(failureTable.failureTime(failureTable.T==300),BinWidth=2) xlabel("Failure Time"); title("Failure Times at T = 300 K")
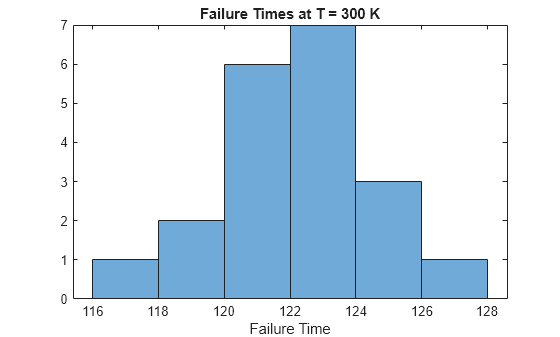
The histogram shows that the failure times at the first stressor level are approximately normally distributed.
Create a plot of mean failure time versus stressor level.
s = groupsummary(failureTable,"T","mean","failureTime"); plot(s.T,s.mean_failureTime,"-o") xlabel("T") ylabel("Failure Time");
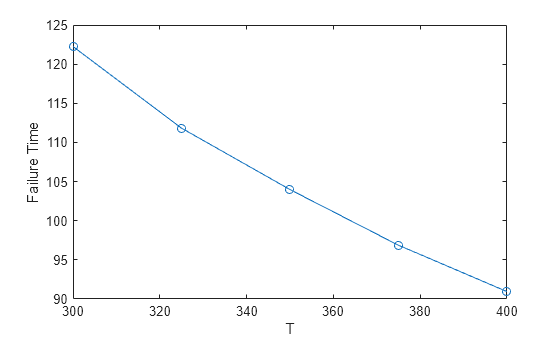
The plot indicates that the mean failure time has a nonlinear dependence on temperature. The failure times in this data set follow an Eyring relationship, which has the functional form f(T) = 1/T*exp(–(b0–b1/T)), where b0 and b1 are model coefficients.
Fit Accelerated Life Model
Fit an accelerated life model to the data using the fitacclife function. Specify a normal life distribution and an Eyring life stress model. Use 230 K as the baseline stressor level.
mdl = fitacclife(failureTable,"failureTime", ... Censoring=failureTable.censored,Distribution="normal", ... StressModel="eyring",StressorName="T",BaselineStressorLevel=230)
mdl =
AcceleratedLifeModel
Life distribution: normal
Stress model: eyring
Baseline stressor level: 230
T NormalMu MeanFailureTime AccelerationFactor
___ ________ _______________ __________________
400 90.742 90.742 1.7712
375 96.951 96.951 1.6577
350 104.07 104.07 1.5443
325 112.32 112.32 1.4309
300 121.99 121.99 1.3175
230 160.72 160.72 1
Log-likelihood: -195.5267
mdl is an AcceleratedLifeModel object, which you can use to compute mean failure times, calculate failure probabilities at specific stressor levels, and create plots. The first column of the output contains the unique stressor levels in the data and the baseline stressor level. The second and third columns list the fitted life distribution parameter values and mean failure times, respectively. The fourth column lists the acceleration factor, which is the ratio of the mean failure time at the stressor level to the mean failure time at the baseline stressor level.
Display information about the fitted model coefficients.
mdl.Coefficients
ans=3×3 table
Source Estimate SE
______________ ________ ________
b0 "StressModel" -10.475 0.016919
b1 "StressModel" 9.8762 5.709
NormalSigma "Distribution" 1.7779 0.13054
The table lists the estimated value and the standard error of each coefficient in the life stress model, and the estimated value and the standard error of the life distribution parameter. The life distribution parameter (NormalSigma) and the first coefficient (b0) of the life stress model are well constrained. The second coefficient of the life stress model (b1) is not well constrained.
List the 95% confidence intervals for each fitted model coefficient and parameter.
coefci(mdl)
ans = 3×2
-10.5084 -10.4412
-1.4545 21.2070
1.5188 2.0370
Display the mean failure times according to the model.
meanfailtime(mdl)
ans=6×2 table
T MeanFailureTime
___ _______________
400 90.742
375 96.951
350 104.07
325 112.32
300 121.99
230 160.72
The last row in the table indicates that the mean failure time at the baseline stressor level (T = 230) is 160.7.
Create a plot of the mean failure times.
meanfailplot(mdl)
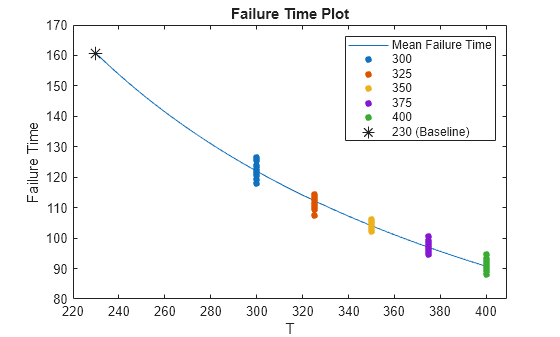
The plot indicates that the Eyring model provides a good fit to the mean failure times at different stressor levels. The mean predicted failure time at the baseline stressor level is indicated on the plot with an asterisk marker.
Create a probability plot of the model and the data.
probplot(mdl)
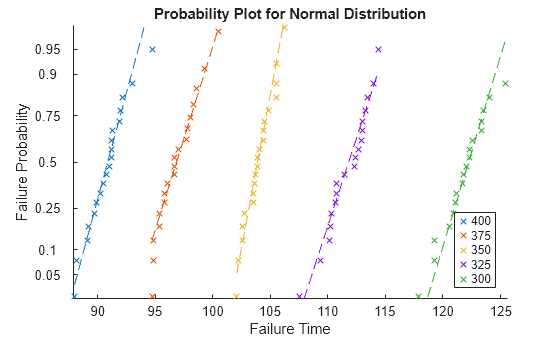
The log-linear plot shows each observation in the data, grouped by stressor level. Each dashed reference line connects the first and third quartiles of the failure time data for that stressor level and extends to the ends of the data. The failure times are shorter at higher temperatures. Also, at a fixed stressor level, the failure probability increases nonlinearly with time.
Load the partFailure data set, which contains simulated observations of failure times for an assembly line part at specific humidity and temperature levels.
load partFailure.matFit an accelerated life model to the data in the partFailure table using the fitacclife function. Use the FailureTime table variable as the failure times, and the other table variables as the stressors.
mdl = fitacclife(partFailure,"FailureTime")mdl =
AcceleratedLifeModel
Life distribution: weibull
Stress model: arrhenius
Humidity Temperature WeibullA MeanFailureTime
________ ___________ ________ _______________
90 35 1.4502 1.3928
90 30 1.5114 1.4516
90 25 1.6015 1.5382
90 20 1.7468 1.6777
90 15 2.0189 1.939
90 12 2.3333 2.241
90 8 3.3507 3.2181
90 5 6.4271 6.1729
80 35 1.4458 1.3886
80 30 1.5069 1.4473
80 25 1.5967 1.5335
80 20 1.7415 1.6727
80 15 2.0128 1.9332
80 12 2.3263 2.2343
80 8 3.3406 3.2084
80 5 6.4077 6.1543
70 35 1.4402 1.3832
70 30 1.501 1.4417
70 25 1.5905 1.5276
70 20 1.7348 1.6662
70 15 2.005 1.9257
70 12 2.3173 2.2256
70 8 3.3276 3.196
70 5 6.3829 6.1304
60 35 1.4328 1.3761
60 30 1.4933 1.4342
60 25 1.5823 1.5197
60 20 1.7259 1.6576
60 15 1.9946 1.9158
60 12 2.3053 2.2141
60 8 3.3104 3.1795
60 5 6.3499 6.0988
50 35 1.4224 1.3662
50 30 1.4825 1.4239
50 25 1.5709 1.5087
50 20 1.7134 1.6456
50 15 1.9803 1.9019
50 12 2.2887 2.1981
50 8 3.2866 3.1566
50 5 6.3041 6.0548
Log-likelihood: 47.0683
mdl is an AcceleratedLifeModel object, which contains information about the fitted model coefficient estimates. By default, the fitacclife function fits an Arrhenius life stress model to the data, and uses a Weibull life distribution. The first and second columns of the displayed output list the unique stressor levels in partFailure. The third and fourth columns list the fitted life distribution parameter values and mean failure times, respectively.
Display information about the fitted model coefficients.
mdl.Coefficients
ans=4×3 table
Source Estimate SE
______________ ________ ________
b0 "StressModel" 1.1592 0.039112
b1 "StressModel" -2.1731 2.0553
b2 "StressModel" 8.6848 0.11616
WeibullB "Distribution" 12.764 0.72011
The table lists the estimated value and the standard error of each coefficient in the life stress model (b0, b1, and b2), and lists the estimated value and the standard error of the life distribution parameter (WeibullB).
List the 95% confidence intervals for each fitted model coefficient and parameter.
ci = coefci(mdl)
ci = 4×2
1.0819 1.2364
-6.2330 1.8868
8.4554 8.9143
11.3419 14.1867
All the coefficients are well constrained except the b1 coefficient of the Arrhenius life stress model.
Load the partFailure data set, which contains simulated observations of failure times for an assembly line part at specific humidity and temperature levels.
load partFailure.matFit an accelerated life model to the data in the partFailure table using the fitacclife function. Use the FailureTime table variable as the failure times, and the other table variables as the stressors. Set the baseline stressor level to a humidity of 50% and a temperature of 20 degrees Celsius.
mdl = fitacclife(partFailure,"FailureTime",BaselineStressorLevel=[50 20]);Display the mean failure times of the model at each stressor level in the data.
meanfailtime(mdl)
ans=41×3 table
Humidity Temperature MeanFailureTime
________ ___________ _______________
90 35 1.3928
90 30 1.4516
90 25 1.5382
90 20 1.6777
90 15 1.939
90 12 2.241
90 8 3.2181
90 5 6.1729
80 35 1.3886
80 30 1.4473
80 25 1.5335
80 20 1.6727
80 15 1.9332
80 12 2.2343
80 8 3.2084
80 5 6.1543
⋮
Create a table of acceleration factors for the unique stressor levels in mdl.
tbl = accelfactor(mdl)
tbl=41×4 table
Humidity Temperature MeanFailureTime AccelerationFactor
________ ___________ _______________ __________________
90 35 1.3928 1.1815
90 30 1.4516 1.1336
90 25 1.5382 1.0699
90 20 1.6777 0.98087
90 15 1.939 0.84869
90 12 2.241 0.73432
90 8 3.2181 0.51136
90 5 6.1729 0.26659
80 35 1.3886 1.1851
80 30 1.4473 1.1371
80 25 1.5335 1.0731
80 20 1.6727 0.98383
80 15 1.9332 0.85125
80 12 2.2343 0.73654
80 8 3.2084 0.51291
80 5 6.1543 0.2674
⋮
Each acceleration factor is the ratio of the mean failure time at the baseline stressor level to the mean failure time at the stressor level.
Create a 3-D scatter plot showing the acceleration factors of different stressor levels.
scatter3(tbl,"Humidity","Temperature","AccelerationFactor")
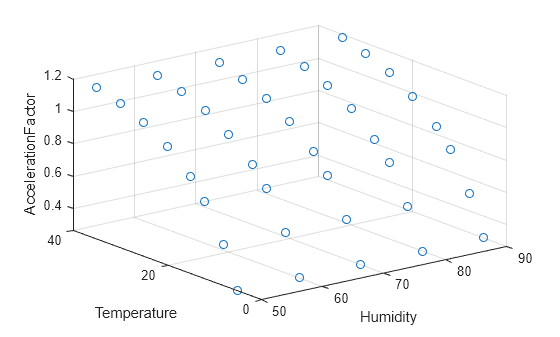
Load the partFailure data set, which contains simulated observations of failure times for an assembly line part at specific humidity and temperature levels.
load partFailure.matFit an accelerated life model to the data in the partFailure table using the fitacclife function. Use the FailureTime table variable as the failure times, and the other table variables as the stressors.
mdl = fitacclife(partFailure,"FailureTime");Plot Survivor Function
Return the survivor function values of the model, evaluated at 50 equally spaced time values between 0.8 and 10. By default, the software calculates the survivor function at the lowest unique stressor level in mdl.StressorLevels. In this example, the lowest unique stressor level corresponds to a humidity value of 50% and a temperature of 5 degrees Celsius.
points = linspace(0.8,10,50)'; x = distfcn(mdl,EvaluationTimes=points)
x = 50×1
0.8000
0.9878
1.1755
1.3633
1.5510
1.7388
1.9265
2.1143
2.3020
2.4898
2.6776
2.8653
3.0531
3.2408
3.4286
⋮
Plot the survivor function at the evaluation points.
distplot(mdl,EvaluationTimes=points)
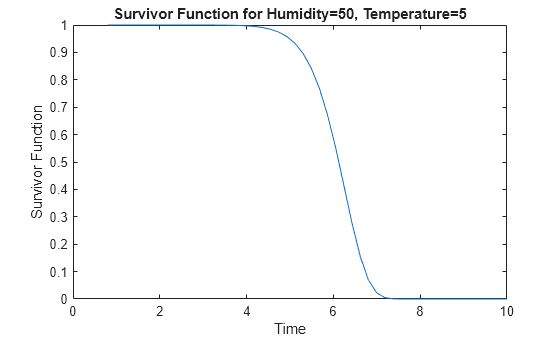
The plot shows that, at this stressor level, the assembly line part has a survival probability close to 100% for time values smaller than 4. The survival probability drops to approximately zero for time values greater than 7.5.
Plot Probability Density Function
Plot the probability density function (pdf).
distplot(mdl,Type="pdf",EvaluationTimes=points)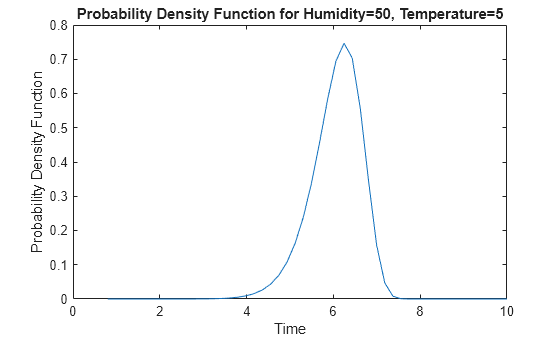
The plot shows that the most likely failure time at this stressor level is approximately 6.2.
Plot Cumulative Distribution Function
Plot the cumulative distribution function (cdf) at a stressor level that corresponds to 90% humidity and a temperature of 20 degrees.
distplot(mdl,[90 20],Type="cdf",EvaluationTimes=points)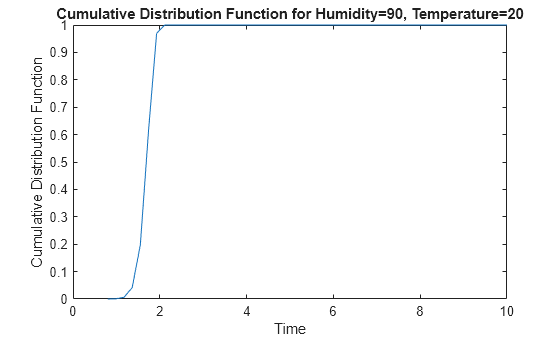
The plot shows that, at this stressor level, approximately half of all assembly line parts have failure times smaller than 1.7.
Plot Inverse Cumulative Distribution Function
Calculate the inverse cumulative distribution function (icdf) at 90% humidity and 35 degrees, and 90% humidity and 20 degrees. Evaluate the icdf at 10 equally spaced probability values between 0.1 and 0.9.
pts = linspace(0.05,0.9,10); icdfval = [icdf(mdl,pts); icdf(mdl,pts,[90,20])]
icdfval = 2×10
4.9953 5.4502 5.6944 5.8737 6.0227 6.1562 6.2834 6.4119 6.5525 6.7298
1.3842 1.5102 1.5779 1.6275 1.6688 1.7058 1.7411 1.7767 1.8156 1.8648
Create a plot of the icdf values for the two stressor levels.
plot(pts,icdfval,"-o") xlabel("Probability") ylabel("ICDF Value") legend(["Humidity = 90, Temperature = 35", ... "Humidity = 90, Temperature = 20"],Location="east")
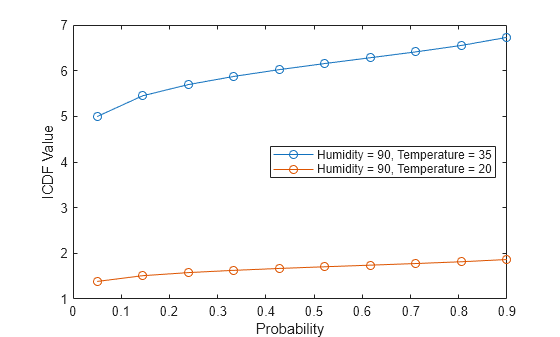
The icdf values are consistently lower at the lower temperature stressor level.
Load the diodeFailure data set, which contains simulated observations of failure times for a diode at different current levels.
load diodeFailure.matFit an accelerated life model to the data in the diodeFailure table using the fitacclife function. Use the FailureTime table variable as the failure times.
mdl = fitacclife(diodeFailure,"FailureTime")mdl =
AcceleratedLifeModel
Life distribution: weibull
Stress model: arrhenius
Current WeibullA MeanFailureTime
_______ ________ _______________
10 1.5323 1.4721
5 2.4956 2.3975
3 4.7821 4.5941
Log-likelihood: 0.4676
mdl is an AcceleratedLifeModel object, which contains information about the fitted model coefficient estimates. By default, the fitacclife function fits an Arrhenius life stress model to the data, using a Weibull life distribution. The first column of the displayed output contains the unique stressor levels in diodeFailure. The second and third columns contain the fitted life distribution coefficient values and mean failure times, respectively.
Create a probability plot.
probplot(mdl)
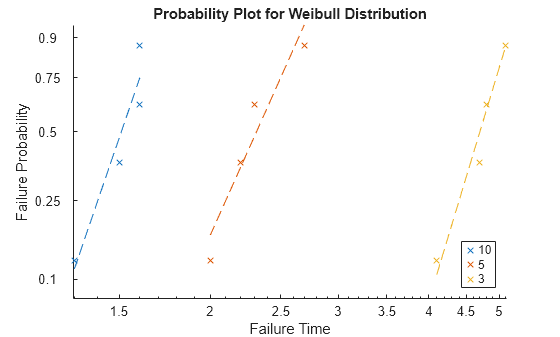
The log-log scatter plot of failure probability versus failure time shows each observation in the data, grouped by stressor level. The dashed lines are linear regression fits to the failure times belonging to each group.
The plot indicates that failure times are shorter at higher current levels, and that the failure probability increases nonlinearly with time at each stressor level.
Load the diodeFailure data set, which contains simulated observations of failure times for a diode at different current levels.
load diodeFailure.matFit an accelerated life model to the data in the diodeFailure table using the fitacclife function. Specify a power law life stress model and use the FailureTime table variable as the failure times.
mdl = fitacclife(diodeFailure,"FailureTime",StressModel="power")
mdl =
AcceleratedLifeModel
Life distribution: weibull
Stress model: power
Current WeibullA MeanFailureTime
_______ ________ _______________
10 1.4869 1.41
5 2.8445 2.6973
3 4.588 4.3506
Log-likelihood: -3.6697
mdl is an AcceleratedLifeModel object, which contains information about the fitted model coefficient estimates. By default, the fitacclife function uses a Weibull life distribution. The first column of the displayed output lists the unique stressor levels in diodeFailure. The second and third columns list the fitted life distribution values and mean failure times, respectively.
Use the meanfailtime function to compute the predicted mean failure time and the Weibull life distribution parameter at current levels 12 and 15.
[meanFailTimes,lifeCoeffs] = meanfailtime(mdl,[12; 15])
meanFailTimes=2×2 table
Current MeanFailureTime
_______ _______________
12 1.1888
15 0.96474
lifeCoeffs=2×2 table
Current WeibullA
_______ ________
12 1.2537
15 1.0174
Create a plot of failure time versus current level.
meanfailplot(mdl)
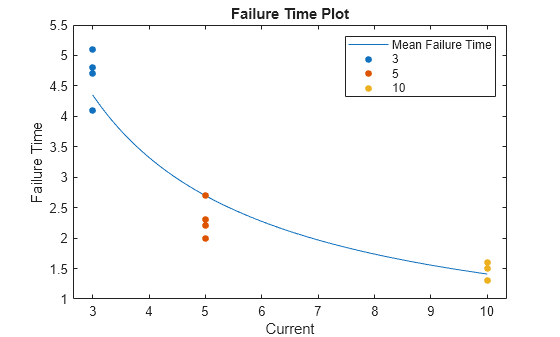
The plot shows that the mean failure time decreases nonlinearly with increasing current level.
More About
A survivor function is the probability of survival as a function of time. It is also called the survival function.
The survivor function gives the probability that the survival time of an individual exceeds a certain value. Because the cumulative distribution function F(t) is the probability that the survival time is less than or equal to a given point t in time, the survivor function for a continuous distribution S(t) is the complement of the cumulative distribution function: S(t) = 1 – F(t).
Version History
Introduced in R2026a
See Also
fitacclife | accelfactor | coefci | distfcn | distplot | icdf | meanfailplot | meanfailtime | probplot
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)