resubPredict
Predict responses for training data using trained regression model
Syntax
Description
yFit = resubPredict(Mdl,Name=Value)IncludeInteractions=true specifies to include interaction terms in
computations for generalized additive models.
[
also returns the standard deviations and prediction intervals of the response variable,
evaluated at each observation in the predictor data yFit,ySD,yInt] = resubPredict(___)Mdl.X, using any of
the input argument combinations in the previous syntaxes. This syntax applies only to
generalized additive models for which IsStandardDeviationFit is true, and to Gaussian process
regression models for which the PredictMethod is not
'bcd'.
Examples
Train a generalized additive model (GAM), then predict responses for the training data.
Load the patients data set.
load patientsCreate a table that contains the predictor variables (Age, Diastolic, Smoker, Weight, Gender, SelfAssessedHealthStatus) and the response variable (Systolic).
tbl = table(Age,Diastolic,Smoker,Weight,Gender,SelfAssessedHealthStatus,Systolic);
Train a univariate GAM that contains the linear terms for the predictors in tbl.
Mdl = fitrgam(tbl,"Systolic")Mdl =
RegressionGAM
PredictorNames: {'Age' 'Diastolic' 'Smoker' 'Weight' 'Gender' 'SelfAssessedHealthStatus'}
ResponseName: 'Systolic'
CategoricalPredictors: [3 5 6]
ResponseTransform: 'none'
Intercept: 122.7800
IsStandardDeviationFit: 0
NumObservations: 100
Properties, Methods
Mdl is a RegressionGAM model object.
Predict responses for the training set.
yFit = resubPredict(Mdl);
Create a table containing the observed response values and the predicted response values. Display the first eight rows of the table.
t = table(tbl.Systolic,yFit, ... 'VariableNames',{'Observed Value','Predicted Value'}); head(t)
Observed Value Predicted Value
______________ _______________
124 124.75
109 109.48
125 122.89
117 115.87
122 121.61
121 122.02
130 126.39
115 115.95
Train a Gaussian process regression (GPR) model by using the fitrgp function. Then predict responses for the training data and estimate prediction intervals of the responses at each observation in the training data by using the resubPredict function.
Generate a training data set.
rng(1) % For reproducibility
n = 100000;
X = linspace(0,1,n)';
X = [X,X.^2];
y = 1 + X*[1;2] + sin(20*X*[1;-2]) + 0.2*randn(n,1);Train a GPR model using the squared exponential kernel function. Estimate parameters by using the subset of regressors ('sr') approximation method, and make predictions using the subset of data ('sd') method. Use 50 points in the active set, and specify 'sgma' (sparse greedy matrix approximation) method for active set selection. Because the scales of the first and second predictors are different, standardize the data set.
gprMdl = fitrgp(X,y,'KernelFunction','squaredExponential', ... 'FitMethod','sr','PredictMethod','sd', ... 'ActiveSetSize',50,'ActiveSetMethod','sgma','Standardize',true);
fitrgp accepts any combination of fitting, prediction, and active set selection methods. However, if you train a model using the block coordinate descent prediction method ('PredictMethod','bcd'), you cannot use the model to compute the standard deviations of the predicted responses; therefore, you also cannot use the model to compute the prediction intervals. For more details, see Tips.
Use the trained model to predict responses for the training data and to estimate the prediction intervals of the predicted responses.
[ypred,~,yci] = resubPredict(gprMdl);
Plot the true responses, predicted responses, and prediction intervals.
figure plot(y,'r') hold on plot(ypred,'b') plot(yci(:,1),'k--') plot(yci(:,2),'k--') legend('True responses','GPR predictions','95% prediction limits','Location','Best') xlabel('X') ylabel('y') hold off
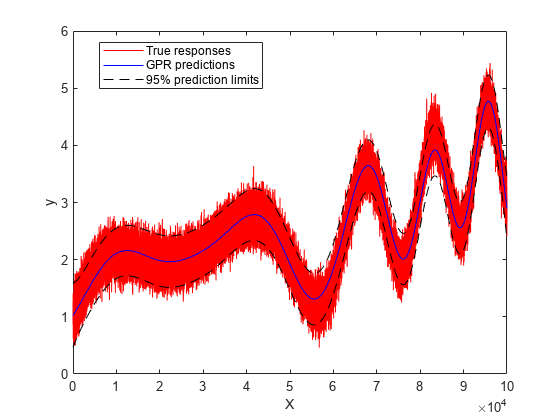
Compute the mean squared error loss on the training data using the trained GPR model.
L = resubLoss(gprMdl)
L = 0.0523
Predict responses for a training data set using a generalized additive model (GAM) that contains both linear and interaction terms for predictors. Specify whether to include interaction terms when predicting responses.
Load the carbig data set, which contains measurements of cars made in the 1970s and early 1980s.
load carbigSpecify Acceleration, Displacement, Horsepower, and Weight as the predictor variables (X) and MPG as the response variable (Y).
X = [Acceleration,Displacement,Horsepower,Weight]; Y = MPG;
Train a generalized additive model that contains all the available linear and interaction terms in X.
Mdl = fitrgam(X,Y,'Interactions','all');
Mdl is a RegressionGAM model object.
Predict the responses using both linear and interaction terms, and then using only linear terms. To exclude interaction terms, specify 'IncludeInteractions',false.
yFit = resubPredict(Mdl);
yFit_nointeraction = resubPredict(Mdl,'IncludeInteractions',false);Create a table containing the observed response values and the predicted response values. Display the first eight rows of the table.
t = table(Mdl.Y,yFit,yFit_nointeraction, ... 'VariableNames',{'Observed Response', ... 'Predicted Response','Predicted Response Without Interactions'}); head(t)
Observed Response Predicted Response Predicted Response Without Interactions
_________________ __________________ _______________________________________
18 18.026 17.22
15 15.003 15.791
18 17.663 16.18
16 16.178 15.536
17 17.107 17.361
15 14.943 14.424
14 14.119 14.981
14 13.864 13.498
Input Arguments
Regression machine learning model, specified as a full regression model object, as given in the following table of supported models.
| Model | Regression Model Object |
|---|---|
| Gaussian process regression model | RegressionGP |
| Generalized additive model (GAM) | RegressionGAM |
| Neural network model | RegressionNeuralNetwork |
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: Alpha=0.01,IncludeInteractions=false specifies the confidence
level as 99% and excludes interaction terms from computations for a generalized additive
model.
Significance level for the confidence level of the prediction intervals
yInt, specified as a numeric scalar in the range
[0,1]. The confidence level of yInt is equal
to 100(1 – Alpha)%.
This argument is valid only for a generalized additive model object that includes the standard deviation fit, or a Gaussian process regression model that does not use the block coordinate descent method for prediction. That is, you can specify this argument only in one of these situations:
MdlisRegressionGAMand theIsStandardDeviationFitproperty ofMdlistrue.MdlisRegressionGPand thePredictMethodproperty ofMdlis not'bcd'.
Example: Alpha=0.01
Data Types: single | double
Flag to include interaction terms of the model, specified as true or
false. This argument is valid only for a generalized
additive model. That is, you can specify this argument only when
Mdl is RegressionGAM.
The default value is true if Mdl contains interaction
terms. The value must be false if the model does not contain interaction
terms.
Data Types: logical
Since R2024b
Output type for the predicted responses yFit, specified as
"matrix" or "table". This argument is valid
only for a neural network model with multiple response variables. That is, you can
specify this argument only when Mdl is a RegressionNeuralNetwork object, where Mdl.Y contains
data for multiple response variables.
Example: OutputType="table"
Data Types: char | string
Since R2023b
Predicted response value to use for observations with missing predictor values,
specified as "median", "mean", or a numeric
scalar. This argument is valid only for a Gaussian process regression or neural
network model. That is, you can specify this argument only when
Mdl is a RegressionGP or RegressionNeuralNetwork object.
| Value | Description |
|---|---|
"median" |
This value is the
default when |
"mean" | resubPredict uses the mean of the observed response
values in the training data as the predicted response value for observations
with missing predictor values. |
| Numeric scalar | resubPredict uses this value as the predicted
response value for observations with missing predictor values. |
Example: PredictionForMissingValue="mean"
Example: PredictionForMissingValue=NaN
Data Types: single | double | char | string
Output Arguments
Predicted responses, returned as a numeric vector, matrix, or table.
If
yFitis a vector, then it has length n, where n is the number of observations in the predictor data (Mdl.X).If
yFitis a matrix or table, then it has n rows, where n is the number of observations in the predictor data.yFitis a matrix or table only whenMdlis a multiresponse regression neural network model.
Standard deviations of the response variable, evaluated at each observation in the
predictor data Mdl.XMdl.Xith element ySD(i) contains the standard deviation
of the ith response for the ith observation
Mdl.X(i,:), estimated using the trained standard deviation model in
Mdl.
This argument is valid only for a generalized additive model object that includes
the standard deviation fit, or a Gaussian process regression model that does not use the
block coordinate descent method for prediction. That is,
resubPredict can return this argument only in one of these situations:
MdlisRegressionGAMand theIsStandardDeviationFitproperty ofMdlistrue.MdlisRegressionGPand thePredictMethodproperty ofMdlis not'bcd'.
Prediction intervals of the response variable, evaluated at each observation in the
predictor data Mdl.XMdl.Xith row yInt(i,:) contains the
100(1 – prediction
interval of the Alpha)%ith response for the ith
observation Mdl.X(i,:). The Alpha value is the
probability that the prediction interval does not contain the true response value
Mdl.Y(i). The first column of yInt contains
the lower limits of the prediction intervals, and the second column contains the upper
limits.
This argument is valid only for a generalized additive model object that includes
the standard deviation fit, or a Gaussian process regression model that does not use the
block coordinate descent method for prediction. That is,
resubPredict can return this argument only in one of these
situations:
MdlisRegressionGAMand theIsStandardDeviationFitproperty ofMdlistrue.MdlisRegressionGPand thePredictMethodproperty ofMdlis not'bcd'.
Algorithms
resubPredict predicts responses according to the corresponding
predict function of the object (Mdl). For a
model-specific description, see the predict function reference pages in
the following table.
| Model | Regression Model Object (Mdl) | predict Object Function |
|---|---|---|
| Gaussian process regression model | RegressionGP | predict |
| Generalized additive model | RegressionGAM | predict |
| Neural network model | RegressionNeuralNetwork | predict |
Alternative Functionality
To compute the predicted responses for new predictor data, use the corresponding
predict function of the object (Mdl).
Extended Capabilities
This function fully supports GPU arrays for RegressionGP and RegressionNeuralNetwork model objects. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2015bresubPredict fully supports GPU arrays for RegressionGP model objects.
You can create a neural network regression model with multiple response variables by
using the fitrnet function.
Regardless of the number of response variables, the function returns a
RegressionNeuralNetwork object. You can use the
resubPredict object function to predict the responses for the
training data.
In the call to resubPredict, you can specify whether to return the
predicted response values as a matrix or table by using the OutputType
name-value argument.
resubPredict fully supports GPU arrays for RegressionNeuralNetwork model objects.
Starting in R2023b, when you predict or compute the loss, some regression models allow you to specify the predicted response value for observations with missing predictor values. Specify the PredictionForMissingValue name-value argument to use a numeric scalar, the training set median, or the training set mean as the predicted value. When computing the loss, you can also specify to omit observations with missing predictor values.
This table lists the object functions that support the
PredictionForMissingValue name-value argument. By default, the
functions use the training set median as the predicted response value for observations with
missing predictor values.
| Model Type | Model Objects | Object Functions |
|---|---|---|
| Gaussian process regression (GPR) model | RegressionGP, CompactRegressionGP | loss, predict, resubLoss, resubPredict |
RegressionPartitionedGP | kfoldLoss, kfoldPredict | |
| Gaussian kernel regression model | RegressionKernel | loss, predict |
RegressionPartitionedKernel | kfoldLoss, kfoldPredict | |
| Linear regression model | RegressionLinear | loss, predict |
RegressionPartitionedLinear | kfoldLoss, kfoldPredict | |
| Neural network regression model | RegressionNeuralNetwork, CompactRegressionNeuralNetwork | loss, predict, resubLoss, resubPredict |
RegressionPartitionedNeuralNetwork | kfoldLoss, kfoldPredict | |
| Support vector machine (SVM) regression model | RegressionSVM, CompactRegressionSVM | loss, predict, resubLoss, resubPredict |
RegressionPartitionedSVM | kfoldLoss, kfoldPredict |
In previous releases, the regression model loss and predict functions listed above used NaN predicted response values for observations with missing predictor values. The software omitted observations with missing predictor values from the resubstitution ("resub") and cross-validation ("kfold") computations for prediction and loss.
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)