predict
Predict labels for Gaussian kernel classification model
Description
Examples
Predict the training set labels using a binary kernel classification model, and display the confusion matrix for the resulting classification.
Load the ionosphere data set. This data set has 34 predictors and 351 binary responses for radar returns, either bad ('b') or good ('g').
load ionosphereTrain a binary kernel classification model that identifies whether the radar return is bad ('b') or good ('g').
rng('default') % For reproducibility Mdl = fitckernel(X,Y);
Mdl is a ClassificationKernel model.
Predict the training set, or resubstitution, labels.
label = predict(Mdl,X);
Construct a confusion matrix.
ConfusionTrain = confusionchart(Y,label);
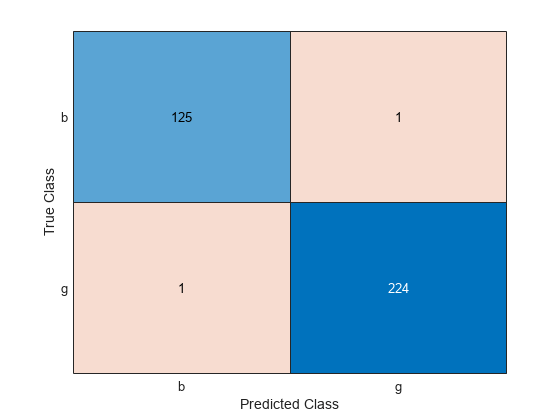
The model misclassifies one radar return for each class.
Predict the test set labels using a binary kernel classification model, and display the confusion matrix for the resulting classification.
Load the ionosphere data set. This data set has 34 predictors and 351 binary responses for radar returns, either bad ('b') or good ('g').
load ionospherePartition the data set into training and test sets. Specify a 15% holdout sample for the test set.
rng('default') % For reproducibility Partition = cvpartition(Y,'Holdout',0.15); trainingInds = training(Partition); % Indices for the training set testInds = test(Partition); % Indices for the test set
Train a binary kernel classification model using the training set. A good practice is to define the class order.
Mdl = fitckernel(X(trainingInds,:),Y(trainingInds),'ClassNames',{'b','g'});
Predict the training-set labels and the test set labels.
labelTrain = predict(Mdl,X(trainingInds,:)); labelTest = predict(Mdl,X(testInds,:));
Construct a confusion matrix for the training set.
ConfusionTrain = confusionchart(Y(trainingInds),labelTrain);
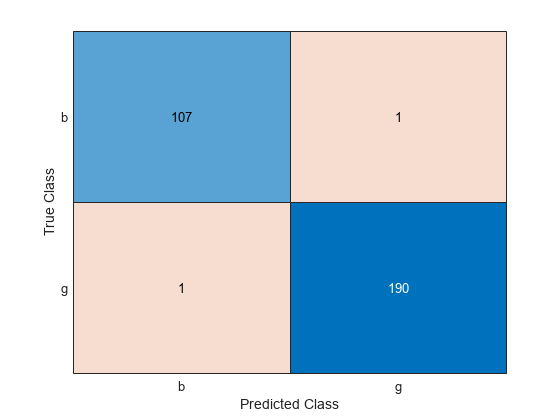
The model misclassifies only one radar return for each class.
Construct a confusion matrix for the test set.
ConfusionTest = confusionchart(Y(testInds),labelTest);
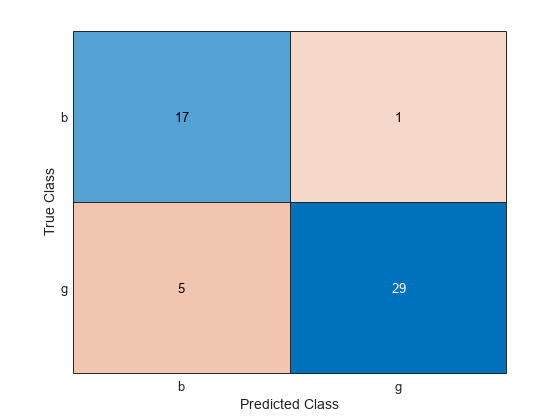
The model misclassifies one bad radar return as being a good return, and five good radar returns as being bad returns.
Estimate posterior class probabilities for a test set, and determine the quality of the model by plotting a receiver operating characteristic (ROC) curve. Kernel classification models return posterior probabilities for logistic regression learners only.
Load the ionosphere data set. This data set has 34 predictors and 351 binary responses for radar returns, either bad ('b') or good ('g').
load ionospherePartition the data set into training and test sets. Specify a 30% holdout sample for the test set.
rng('default') % For reproducibility Partition = cvpartition(Y,'Holdout',0.30); trainingInds = training(Partition); % Indices for the training set testInds = test(Partition); % Indices for the test set
Train a binary kernel classification model. Fit logistic regression learners.
Mdl = fitckernel(X(trainingInds,:),Y(trainingInds), ... 'ClassNames',{'b','g'},'Learner','logistic');
Predict the posterior class probabilities for the test set.
[~,posterior] = predict(Mdl,X(testInds,:));
Because Mdl has one regularization strength, the output posterior is a matrix with two columns and rows equal to the number of test-set observations. Column i contains posterior probabilities of Mdl.ClassNames(i) given a particular observation.
Compute the performance metrics (true positive rates and false positive rates) for a ROC curve and find the area under the ROC curve (AUC) value by creating a rocmetrics object.
rocObj = rocmetrics(Y(testInds),posterior,Mdl.ClassNames);
Plot the ROC curve for the second class by using the plot function of rocmetrics.
plot(rocObj,ClassNames=Mdl.ClassNames(2))
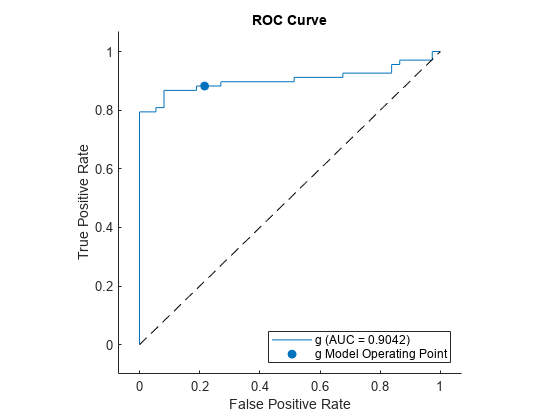
The AUC is close to 1, which indicates that the model predicts labels well.
Input Arguments
Binary kernel classification model, specified as a ClassificationKernel model object. You can create a
ClassificationKernel model object using fitckernel.
Predictor data to be classified, specified as a numeric matrix or table.
Each row of X corresponds to one observation, and
each column corresponds to one variable.
For a numeric matrix:
The variables in the columns of
Xmust have the same order as the predictor variables that trainedMdl.If you trained
Mdlusing a table (for example,Tbl) andTblcontains all numeric predictor variables, thenXcan be a numeric matrix. To treat numeric predictors inTblas categorical during training, identify categorical predictors by using theCategoricalPredictorsname-value pair argument offitckernel. IfTblcontains heterogeneous predictor variables (for example, numeric and categorical data types) andXis a numeric matrix, thenpredictthrows an error.
For a table:
predictdoes not support multicolumn variables or cell arrays other than cell arrays of character vectors.If you trained
Mdlusing a table (for example,Tbl), then all predictor variables inXmust have the same variable names and data types as those that trainedMdl(stored inMdl.PredictorNames). However, the column order ofXdoes not need to correspond to the column order ofTbl. Also,TblandXcan contain additional variables (response variables, observation weights, and so on), butpredictignores them.If you trained
Mdlusing a numeric matrix, then the predictor names inMdl.PredictorNamesand corresponding predictor variable names inXmust be the same. To specify predictor names during training, see thePredictorNamesname-value pair argument offitckernel. All predictor variables inXmust be numeric vectors.Xcan contain additional variables (response variables, observation weights, and so on), butpredictignores them.
Data Types: table | double | single
Output Arguments
Predicted class labels, returned as a categorical or character array, logical or numeric matrix, or cell array of character vectors.
Label has n rows, where
n is the number of observations in
X, and has the same data type as the observed class
labels (Y) used to train Mdl.
(The software treats string arrays as cell arrays of character
vectors.)
The predict function classifies an observation into the class yielding the highest score. For an observation with NaN scores, the
function classifies the observation into the majority class, which makes up the largest
proportion of the training labels.
Classification scores, returned as an n-by-2
numeric array, where n is the number of observations in
X.
Score(
is the score for classifying observation i,j)i into
class j. Mdl.ClassNames stores
the order of the classes.
If Mdl.Learner is 'logistic', then
classification scores are posterior probabilities.
More About
For kernel classification models, the raw classification score for classifying the observation x, a row vector, into the positive class is defined by
is a transformation of an observation for feature expansion.
β is the estimated column vector of coefficients.
b is the estimated scalar bias.
The raw classification score for classifying x into the negative class is −f(x). The software classifies observations into the class that yields a positive score.
If the kernel classification model consists of logistic regression learners, then the
software applies the 'logit' score transformation to the raw
classification scores (see ScoreTransform).
Extended Capabilities
The
predict function supports tall arrays with the following usage
notes and limitations:
predictdoes not support talltabledata.
For more information, see Tall Arrays.
Usage notes and limitations:
Use
saveLearnerForCoder,loadLearnerForCoder, andcodegen(MATLAB Coder) to generate code for thepredictfunction. Save a trained model by usingsaveLearnerForCoder. Define an entry-point function that loads the saved model by usingloadLearnerForCoderand calls thepredictfunction. Then usecodegento generate code for the entry-point function.To generate single-precision C/C++ code for
predict, specifyDataType="single"when you call theloadLearnerForCoderfunction.If the code generator uses the Open Multiprocessing (OpenMP) library, the generated code of
predictsplits the predictor dataXinto multiple chunks and predicts responses for the chunks in parallel. The generated code usesparfor(MATLAB Coder) to create loops that run in parallel on supported shared-memory multicore platforms. If your compiler does not support the OpenMP application interface, or if you disable the OpenMP library, the generated code does not split the predictor data and, therefore, processes one observation at a time. To find supported compilers, see Supported Compilers. To disable the OpenMP library, set theEnableOpenMPproperty of the configuration object tofalse. For details, seecoder.CodeConfig(MATLAB Coder).This table contains notes about the arguments of
predict. Arguments not included in this table are fully supported.Argument Notes and Limitations MdlFor the usage notes and limitations of the model object, see Code Generation of the
ClassificationKernelobject.XFor general code generation,
Xmust be a single-precision or double-precision matrix or a table containing numeric variables, categorical variables, or both.The number of rows, or observations, in
Xcan be a variable size, but the number of columns inXmust be fixed.If you want to specify
Xas a table, then your model must be trained using a table, and your entry-point function for prediction must do the following:Accept data as arrays.
Create a table from the data input arguments and specify the variable names in the table.
Pass the table to
predict.
For an example of this table workflow, see Generate Code to Classify Data in Table. For more information on using tables in code generation, see Code Generation for Tables (MATLAB Coder) and Table Limitations for Code Generation (MATLAB Coder).
For more information, see Introduction to Code Generation for Statistics and Machine Learning Functions.
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2017bpredict fully supports GPU arrays.
You can generate C/C++ code for the predict function.
See Also
ClassificationKernel | fitckernel | resume | rocmetrics | confusionchart
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)