Work with Whole Slide Images Using OpenSlide and Bio-Formats
Import whole slide imaging (WSI) files using the Medical Imaging Toolbox™ Interface for Whole Slide Imaging File Reader support package, which reads whole slide images using the OpenSlide and Bio-Formats libraries. You can install the support package from Add-On Explorer. For more information about installing add-ons, see Get and Manage Add-Ons.
Whole slide images are high-resolution digital images of microscope slides used in digital pathology. Whole slide images can be challenging to work with, because they come in many proprietary formats that require specialized file readers. Additionally, WSI files are large and typically contain multiresolution image pyramids, associated images such as low-resolution thumbnails or macro images, and rich metadata.
To address these challenges, the support package imports whole slide images as
blockedImage objects, which are designed to work with large,
multiresolution images and nonstandard image formats. Instead of storing an image as a single
array, a blockedImage object represents each resolution level as a collection
of image blocks. Because MATLAB® functions read and process the blocks in a blockedImage object
independently, they load data into memory only when needed. Therefore, blocked images enable
efficient memory usage, parallel processing, and smooth zoom and pan interactions within image
displays.
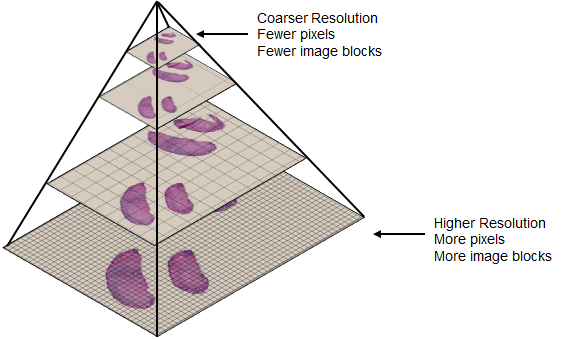
This image illustrates the structure of a tiled, multiresolution whole slide image pyramid read as a blocked image [1]. The finer resolution level at the bottom of the pyramid contains more pixels and requires more blocks to represent, while the top of the pyramid corresponds to a low-resolution thumbnail image.
Choose Between OpenSlide and Bio-Formats Libraries
The functions you call determine which library the support package uses to import your whole slide images. The best library to use depends on your data and preferences. OpenSlide provides a simple, fast interface for standard 2-D digital pathology images. Bio-Formats is more complex and flexible in the number of formats and image types it supports. If both libraries support your data, consider which library is more commonly used within your field.
This table compares the OpenSlide and Bio-Formats libraries used in the support package.
| Library | Functions | Supported File Formats | Supported Image Content |
|---|---|---|---|
| OpenSlide — Quickly read whole slide image files from common digital pathology scanners. |
| Smaller set of formats, such as Aperio (.svs), Hamamatsu
(.vms,.ndpi), Leica
(.scn), and ZEISS (.czi). For a complete
list, see the OpenSlide
Documentation. |
|
| Bio-Formats — Read a broad range of whole slide images and other life sciences images. |
| Over 100 formats listed in the Bio-Formats Documentation, including most of the formats supported by OpenSlide. | In addition to the images supported by OpenSlide:
|
Note
Reading the same file using the OpenSlide and Bio-Formats libraries can yield different results, depending on the proprietary file format and image content. To learn more about how each library interprets image data, including edge cases for specific formats, see the Bio-Formats Library and OpenSlide Library documentation.
Import WSI Metadata and Image Data
This section shows how to read and display WSI data using the support package. The code in this example uses the OpenSlide library. For additional examples showing how to import files using the Bio-Formats library, see bioformatsinfo and bioformatsread.
Download WSI Data
Run this code to download a whole slide image from the MathWorks® website, and unzip the downloaded folder. The image, stored in the NDPI format, contains an H&E stained microscopy slide provided by the OpenSlide library test data set [1].
zipFile = matlab.internal.examples.downloadSupportFile("image","data/CMU-1.zip"); filepath = fileparts(zipFile); if ~exist(filepath) unzip(zipFile,filepath) end filename = fullfile(filepath,"CMU-1.ndpi");
Read Whole Slide Image Metadata
Read the metadata from the file.
info = openslideinfo(filename);
Check the AssociatedImages field. The file contains a macro image, which provides a low-resolution overview of the full slide.
info.AssociatedImages
ans = "macro"
Read Whole Slide Image Data
Read the main image data.
dataIm = openslideread(filename);
Read the macro image data by specifying the ImageType name-value argument as "macro".
macroIm = openslideread(filename,ImageType="macro");Display and Explore Whole Slide Image
You can display the whole slide image by using the imageshow function, which provides fast performance and smooth zoom and pan interactions for large and multiresolution images. The function picks the resolution level to display based on the display size, and the resolution level changes as you zoom in or out. You can customize the image display to add a scale bar, measure distances between points in the image, or label regions of interest using various annotation shapes. To learn more about displaying blocked images using imageshow, see Display and Explore Blocked Images.
imageshow(dataIm)
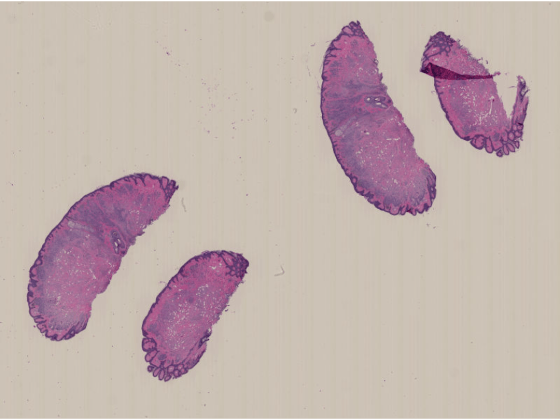
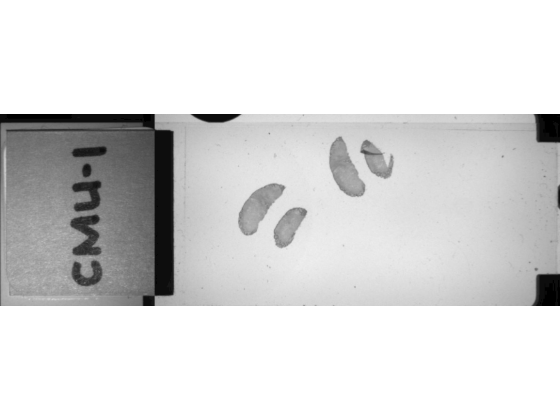
Display the macro images.
imageshow(macroIm)
Process Whole Slide Images for Downstream Tasks
After you import your whole slide image as a blockedImage object, you
can perform additional preprocessing, or perform downstream tasks like image segmentation or
classification. For example, you can:
Extract specific regions using block or image coordinates by using the
getBlockandgetRegionobject functions, or extract specific resolution levels by using thegatherfunction.Apply built-in or custom image preprocessing functions in a blockwise manner by using the
applyobject function. For examples, see Process Blocked Images Efficiently Using Partial Images or Lower Resolutions and Process Blocked Images Efficiently Using Mask.Segment nuclei or cells using Cellpose, as shown in the Detect Nuclei in Large Whole Slide Images Using Cellpose example.
Classify images, such as either healthy or cancerous, as shown in the Classify Tumors in Digital Pathology Images Using Cell Nucleus Morphology example.
References
[1] "CMU‑1.ndpi". OpenSlide Test Data. Accessed November 10, 2025. https://openslide.cs.cmu.edu/download/openslide-testdata/Hamamatsu/.
See Also
bioformatsinfo | bioformatsread | openslideinfo | openslideread | blockedImage | cellpose | segmentCells2D | imageshow
Topics
- Display and Explore Blocked Images
- Detect Nuclei in Large Whole Slide Images Using Cellpose
- Read Whole-Slide Images with OpenSlide