edge
Classification edge for neural network classifier
Description
e = edge(Mdl,Tbl,ResponseVarName)Mdl using the predictor
data in table Tbl and the class labels in the
ResponseVarName table variable.
e is returned as a scalar value that represents the mean of the
classification margins.
e = edge(___,Name,Value)
Note
If the predictor data X or the predictor variables in
Tbl contain any missing values, the
edge function can return NaN. For more
details, see edge can return NaN for predictor data with missing values.
Examples
Calculate the test set classification edge of a neural network classifier.
Load the patients data set. Create a table from the data set. Each row corresponds to one patient, and each column corresponds to a diagnostic variable. Use the Smoker variable as the response variable, and the rest of the variables as predictors.
load patients
tbl = table(Diastolic,Systolic,Gender,Height,Weight,Age,Smoker);Separate the data into a training set tblTrain and a test set tblTest by using a stratified holdout partition. The software reserves approximately 30% of the observations for the test data set and uses the rest of the observations for the training data set.
rng("default") % For reproducibility of the partition c = cvpartition(tbl.Smoker,"Holdout",0.30); trainingIndices = training(c); testIndices = test(c); tblTrain = tbl(trainingIndices,:); tblTest = tbl(testIndices,:);
Train a neural network classifier using the training set. Specify the Smoker column of tblTrain as the response variable. Specify to standardize the numeric predictors.
Mdl = fitcnet(tblTrain,"Smoker", ... "Standardize",true);
Calculate the test set classification edge.
e = edge(Mdl,tblTest,"Smoker")e = 0.8657
The mean of the classification margins is close to 1, which indicates that the model performs well overall.
Perform feature selection by comparing test set classification margins, edges, errors, and predictions. Compare the test set metrics for a model trained using all the predictors to the test set metrics for a model trained using only a subset of the predictors.
Load the sample file fisheriris.csv, which contains iris data including sepal length, sepal width, petal length, petal width, and species type. Read the file into a table.
fishertable = readtable('fisheriris.csv');Separate the data into a training set trainTbl and a test set testTbl by using a stratified holdout partition. The software reserves approximately 30% of the observations for the test data set and uses the rest of the observations for the training data set.
rng("default") c = cvpartition(fishertable.Species,"Holdout",0.3); trainTbl = fishertable(training(c),:); testTbl = fishertable(test(c),:);
Train one neural network classifier using all the predictors in the training set, and train another classifier using all the predictors except PetalWidth. For both models, specify Species as the response variable, and standardize the predictors.
allMdl = fitcnet(trainTbl,"Species","Standardize",true); subsetMdl = fitcnet(trainTbl,"Species ~ SepalLength + SepalWidth + PetalLength", ... "Standardize",true);
Calculate the test set classification margins for the two models. Because the test set includes only 45 observations, display the margins using bar graphs.
For each observation, the classification margin is the difference between the classification score for the true class and the maximal score for the false classes. Because neural network classifiers return classification scores that are posterior probabilities, margin values close to 1 indicate confident classifications and negative margin values indicate misclassifications.
tiledlayout(2,1) % Top axes ax1 = nexttile; allMargins = margin(allMdl,testTbl); bar(ax1,allMargins) xlabel(ax1,"Observation") ylabel(ax1,"Margin") title(ax1,"All Predictors") % Bottom axes ax2 = nexttile; subsetMargins = margin(subsetMdl,testTbl); bar(ax2,subsetMargins) xlabel(ax2,"Observation") ylabel(ax2,"Margin") title(ax2,"Subset of Predictors")

Compare the test set classification edge, or mean of the classification margins, of the two models.
allEdge = edge(allMdl,testTbl)
allEdge = 0.8198
subsetEdge = edge(subsetMdl,testTbl)
subsetEdge = 0.9556
Based on the test set classification margins and edges, the model trained on a subset of the predictors seems to outperform the model trained on all the predictors.
Compare the test set classification error of the two models.
allError = loss(allMdl,testTbl); allAccuracy = 1-allError
allAccuracy = 0.9111
subsetError = loss(subsetMdl,testTbl); subsetAccuracy = 1-subsetError
subsetAccuracy = 0.9778
Again, the model trained using only a subset of the predictors seems to perform better than the model trained using all the predictors.
Visualize the test set classification results using confusion matrices.
allLabels = predict(allMdl,testTbl);
figure
confusionchart(testTbl.Species,allLabels)
title("All Predictors")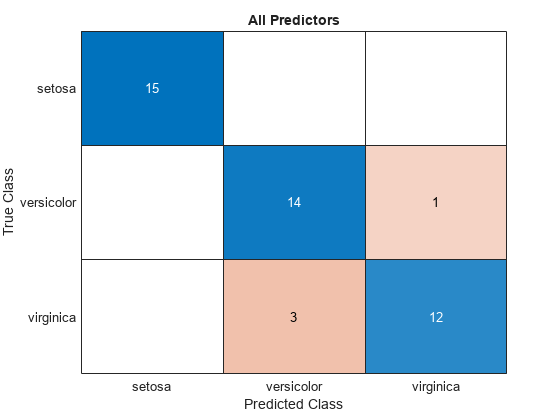
subsetLabels = predict(subsetMdl,testTbl);
figure
confusionchart(testTbl.Species,subsetLabels)
title("Subset of Predictors")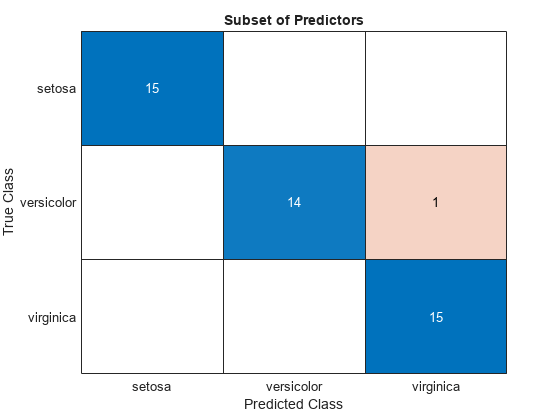
The model trained using all the predictors misclassifies four of the test set observations. The model trained using a subset of the predictors misclassifies only one of the test set observations.
Given the test set performance of the two models, consider using the model trained using all the predictors except PetalWidth.
Input Arguments
Trained neural network classifier, specified as a ClassificationNeuralNetwork model object or CompactClassificationNeuralNetwork model object returned by fitcnet or
compact,
respectively.
Sample data, specified as a table. Each row of Tbl corresponds to one observation, and each column corresponds to one predictor variable. Optionally, Tbl can contain an additional column for the response variable. Tbl must contain all of the predictors used to train Mdl. Multicolumn variables and cell arrays other than cell arrays of character vectors are not allowed.
If
Tblcontains the response variable used to trainMdl, then you do not need to specifyResponseVarNameorY.If you trained
Mdlusing sample data contained in a table, then the input data foredgemust also be in a table.If you set
'Standardize',trueinfitcnetwhen trainingMdl, then the software standardizes the numeric columns of the predictor data using the corresponding means and standard deviations.
Data Types: table
Response variable name, specified as the name of a variable in Tbl. If Tbl contains the response variable used to train Mdl, then you do not need to specify ResponseVarName.
If you specify ResponseVarName, then you must specify it as a character
vector or string scalar. For example, if the response variable is stored as
Tbl.Y, then specify ResponseVarName as
'Y'. Otherwise, the software treats all columns of
Tbl, including Tbl.Y, as predictors.
The response variable must be a categorical, character, or string array; a logical or numeric vector; or a cell array of character vectors. If the response variable is a character array, then each element must correspond to one row of the array.
Data Types: char | string
Class labels, specified as a categorical, character, or string array; logical or numeric vector; or cell array of character vectors.
The data type of
Ymust be the same as the data type ofMdl.ClassNames. (The software treats string arrays as cell arrays of character vectors.)The distinct classes in
Ymust be a subset ofMdl.ClassNames.If
Yis a character array, then each element must correspond to one row of the array.The length of
Ymust be equal to the number of observations inXorTbl.
Data Types: categorical | char | string | logical | single | double | cell
Predictor data, specified as a numeric matrix. By default,
edge assumes that each row of X
corresponds to one observation, and each column corresponds to one predictor
variable.
Note
If you orient your predictor matrix so that observations correspond to columns and
specify 'ObservationsIn','columns', then you might experience a
significant reduction in computation time.
The length of Y and the number of observations in X must be equal.
If you set 'Standardize',true in fitcnet when training Mdl, then the software standardizes the numeric columns of the predictor data using the corresponding means and standard deviations.
Data Types: single | double
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: edge(Mdl,Tbl,"Response","Weights","W") specifies to use the
Response and W variables in the table
Tbl as the class labels and observation weights,
respectively.
Predictor data observation dimension, specified as 'rows' or
'columns'.
Note
If you orient your predictor matrix so that observations correspond to columns and
specify 'ObservationsIn','columns', then you might experience a
significant reduction in computation time. You cannot specify
'ObservationsIn','columns' for predictor data in a
table.
Data Types: char | string
Observation weights, specified as a nonnegative numeric vector or the name of a
variable in Tbl. The software weights each observation in
X or Tbl with the corresponding value in
Weights. The length of Weights must equal
the number of observations in X or
Tbl.
If you specify the input data as a table Tbl, then
Weights can be the name of a variable in
Tbl that contains a numeric vector. In this case, you must
specify Weights as a character vector or string scalar. For
example, if the weights vector W is stored as
Tbl.W, then specify it as 'W'.
By default, Weights is ones(n,1), where
n is the number of observations in X or
Tbl.
If you supply weights, then edge computes the weighted
classification edge and normalizes weights to sum to the value of the prior
probability in the respective class.
Data Types: single | double | char | string
More About
The classification edge is the mean of the
classification margins, or the weighted mean of the
classification margins when you specify
Weights.
One way to choose among multiple classifiers, for example to perform feature selection, is to choose the classifier that yields the greatest edge.
The classification margin for binary classification is, for each observation, the difference between the classification score for the true class and the classification score for the false class. The classification margin for multiclass classification is the difference between the classification score for the true class and the maximal score for the false classes.
If the margins are on the same scale (that is, the score values are based on the same score transformation), then they serve as a classification confidence measure. Among multiple classifiers, those that yield greater margins are better.
Extended Capabilities
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2021aedge fully supports GPU arrays.
The edge function no longer omits an observation with a
NaN score when computing the weighted mean of the classification margins. Therefore,
edge can now return NaN when the predictor data
X or the predictor variables in Tbl
contain any missing values. In most cases, if the test set observations do not contain
missing predictors, the edge function does not return
NaN.
This change improves the automatic selection of a classification model when you use
fitcauto.
Before this change, the software might select a model (expected to best classify new
data) with few non-NaN predictors.
If edge in your code returns NaN, you can update your code
to avoid this result. Remove or replace the missing values by using rmmissing or fillmissing, respectively.
The following table shows the classification models for which the
edge object function might return NaN. For more details,
see the Compatibility Considerations for each edge
function.
| Model Type | Full or Compact Model Object | edge Object Function |
|---|---|---|
| Discriminant analysis classification model | ClassificationDiscriminant, CompactClassificationDiscriminant | edge |
| Ensemble of learners for classification | ClassificationEnsemble, CompactClassificationEnsemble | edge |
| Gaussian kernel classification model | ClassificationKernel | edge |
| k-nearest neighbor classification model | ClassificationKNN | edge |
| Linear classification model | ClassificationLinear | edge |
| Neural network classification model | ClassificationNeuralNetwork, CompactClassificationNeuralNetwork | edge |
| Support vector machine (SVM) classification model | edge |
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)