Signal Labeler
Label signal attributes, regions, and points of interest
Description
The Signal Labeler app is an interactive tool that enables you to label signals for analysis or for use in machine learning and deep learning applications. Using Signal Labeler, you can:
Label signal attributes, regions, and points of interest
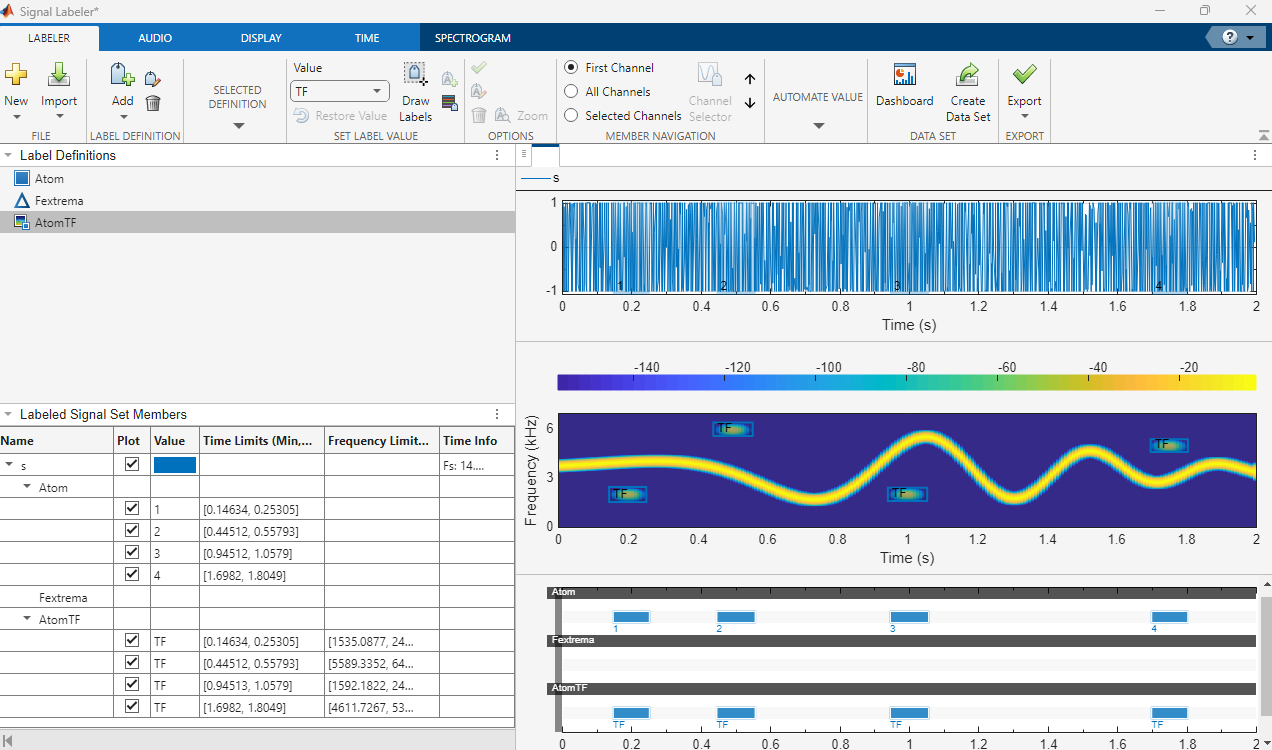
Label spectrogram time-frequency regions of interest (since R2025a)
Use logical, categorical, numerical, or string-valued labels
Automatically label signal peaks or bounded signal regions
Apply custom labeling functions
Import, label, and play audio signals
Use frequency and time-frequency views to aid labeling in time domain
Use time view to aid labeling in time-frequency domain (since R2025a)
Create data sets that include signals, spectrograms, and label masks (since R2025a)
Add, edit, and delete labels or sublabels
Display selected subsets of signals and labels
Signal Labeler saves data as labeledSignalSet
objects. You can export labeledSignalSet objects to MATLAB® or Diagnostic Feature
Designer (Predictive Maintenance Toolbox). You can create data sets as files to train a network, classifier, or analyze
data and report statistics.
For more information, see Use Signal Labeler App.
You need an Audio Toolbox™ license to import, play, and label audio signals.
Audio playback is not supported in MATLAB Online.
You need a Predictive Maintenance Toolbox™ license to export labeled signal sets to Diagnostic Feature Designer.
Signal Labeler no longer supports the feature extraction mode, which is now available as an app. To extract signal features, open Signal Feature Extractor from the MATLAB Toolstrip or the Command Window.
Open the Signal Labeler App
MATLAB Toolstrip: On the Apps tab, under Signal Processing and Audio, click the app icon.
MATLAB command prompt: Enter
signalLabeler.
Examples
Related Examples
- Label Signal Attributes, Regions of Interest, and Points
- Examine Labeled Signal Set
- Automate Signal Labeling with Custom Functions
- Label Spoken Words in Audio Signals
- Label Radar Signals with Signal Labeler (Radar Toolbox)
- Export Labeled Data from Signal Labeler for AI-Based Spectrum Sensing Applications
Programmatic Use
Version History
Introduced in R2019aSee Also
Apps
Functions
Topics
- Use Signal Labeler App
- Import Data into Signal Labeler
- Import and Play Audio File Data in Signal Labeler
- Create or Import Signal Label Definitions
- Label Signals Interactively or Automatically
- Custom Labeling Functions
- Customize Labeling View
- Spectrogram Computation in Signal Labeler
- Dashboard
- Export Data and Create Data Sets
- Signal Labeler Usage Tips