Train Naive Bayes Classifiers Using Classification Learner App
This example shows how to create and compare different naive Bayes classifiers using the Classification Learner app, and export trained models to the workspace to make predictions for new data.
Naive Bayes classifiers leverage Bayes' theorem and make the assumption that predictors are independent of one another within each class. However, the classifiers appear to work well even when the independence assumption is not valid. You can use naive Bayes with two or more classes in Classification Learner. The app allows you to train a Gaussian naive Bayes model or a kernel naive Bayes model individually or simultaneously.
This table lists the available naive Bayes models in Classification Learner and the probability distributions used by each model to fit predictors.
| Model | Numerical Predictor | Categorical Predictor |
|---|---|---|
| Gaussian naive Bayes | Gaussian distribution (or normal distribution) | multivariate multinomial distribution |
| Kernel naive Bayes | Kernel distribution You can specify the kernel type and support. Classification Learner automatically determines the kernel width using the underlying fitcnb
function. | multivariate multinomial distribution |
This example uses Fisher's iris data set, which contains measurements of flowers (petal length, petal width, sepal length, and sepal width) for specimens from three species. Train naive Bayes classifiers to predict the species based on the predictor measurements.
In the MATLAB® Command Window, load the Fisher iris data set and create a table of measurement predictors (or features) using variables from the data set.
fishertable = readtable("fisheriris.csv");Click the Apps tab, and then click the arrow at the right of the Apps section to open the apps gallery. In the Machine Learning and Deep Learning group, click Classification Learner.
On the Learn tab, in the File section, select New Session > From Workspace.

In the New Session from Workspace dialog box, select the table
fishertablefrom the Data Set Variable list (if necessary).As shown in the dialog box, the app selects the response and predictor variables based on their data type. Petal and sepal length and width are predictors, and species is the response that you want to classify. For this example, do not change the selections.

To accept the default validation scheme and continue, click Start Session. The default validation option is cross-validation, to protect against overfitting.
Classification Learner creates a scatter plot of the data.

Use the scatter plot to investigate which variables are useful for predicting the response. Select different options on the X and Y lists under Predictors to visualize the distribution of species and measurements. Observe which variables separate the species colors most clearly.
The
setosaspecies (blue points) is easy to separate from the other two species with all four predictors. Theversicolorandvirginicaspecies are much closer together in all predictor measurements and overlap, especially when you plot sepal length and width.setosais easier to predict than the other two species.Create a naive Bayes model. On the Learn tab, in the Models section, click the arrow to open the gallery. In the Naive Bayes Classifiers group, click Gaussian Naive Bayes. Note that the Model Hyperparameters section of the model Summary tab contains no hyperparameter options.
In the Train section, click Train All and select Train Selected.
Note
If you have Parallel Computing Toolbox™, then the Use Parallel button is selected by default. After you click Train All and select Train All or Train Selected, the app opens a parallel pool of workers. During this time, you cannot interact with the software. After the pool opens, you can continue to interact with the app while models train in parallel.
If you do not have Parallel Computing Toolbox, then the Use Background Training check box in the Train All menu is selected by default. After you select an option to train models, the app opens a background pool. After the pool opens, you can continue to interact with the app while models train in the background.
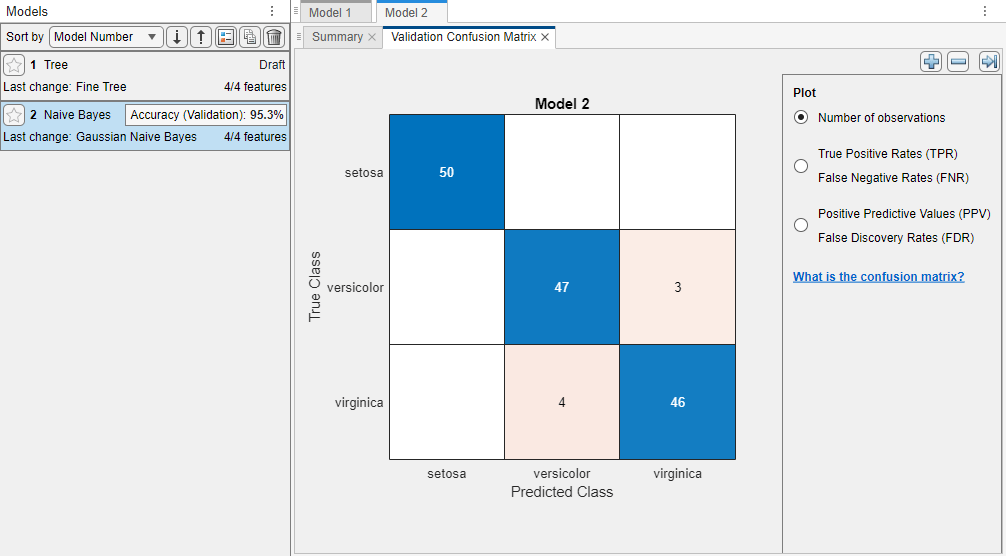
The app creates a Gaussian naive Bayes model, and plots a validation confusion matrix.
The app displays the Gaussian Naive Bayes model in the Models pane. Check the model validation accuracy in the Accuracy (Validation) box. The value shows that the model performs well.
For the Gaussian Naive Bayes model, by default, the app models the distribution of numerical predictors using the Gaussian distribution, and models the distribution of categorical predictors using the multivariate multinomial distribution (MVMN).
Note
Validation introduces some randomness into the results. Your model validation results might vary from the results shown in this example.
Examine the scatter plot for the trained model. On the Learn tab, in the Plots and Results section, click the arrow to open the gallery, and then click Scatter in the Validation Results group. An X indicates a misclassified point. The blue points (
setosaspecies) are all correctly classified, but the other two species have misclassified points. Under Plot, switch between the Data and Model predictions options. Observe the color of the incorrect (X) points. Or, to view only the incorrect points, clear the Correct check box.Train a kernel naive Bayes model for comparison. On the Learn tab, in the Models gallery, click Kernel Naive Bayes. The app displays a draft kernel naive Bayes model in the Models pane.
In the model Summary tab, under Model Hyperparameters, select
Trianglefrom the Kernel type list, selectPositivefrom the Support list, and selectNofrom the Standardize data list.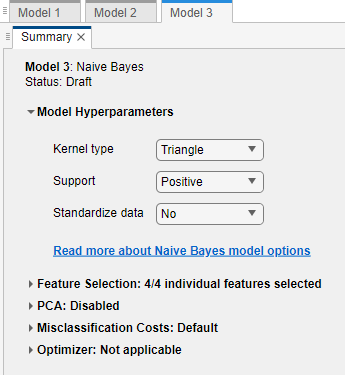
In the Train section, click Train All and select Train Selected to train the new model.
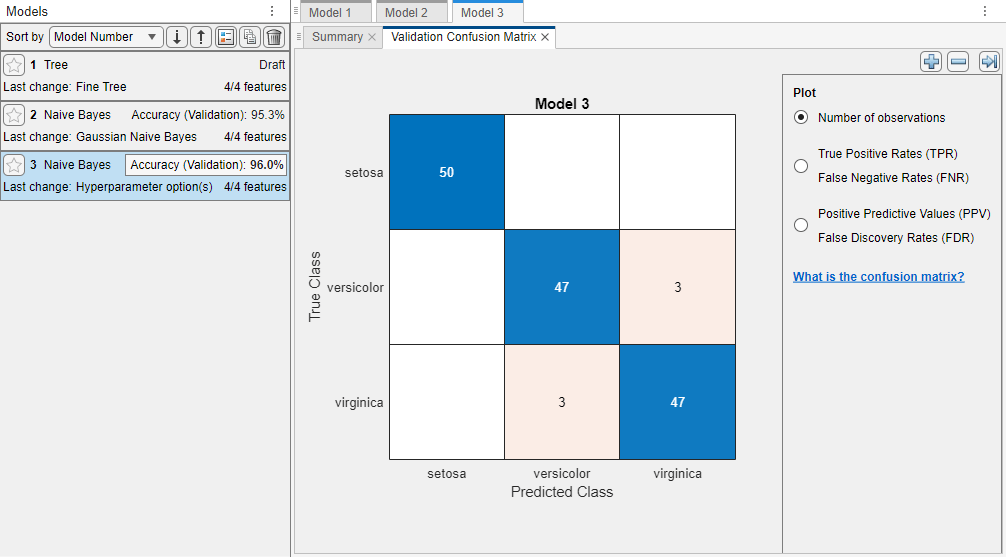
The Models pane displays the model validation accuracy for the new kernel naive Bayes model. Its validation accuracy is better than the validation accuracy of the Gaussian naive Bayes model. The app highlights the Accuracy (Validation) value of the best model (or models) by outlining it in a box.
In the Models pane, click each model to view and compare the results. To view the results for a model, inspect the model Summary tab. The Summary tab displays the Training Results and Additional Training Results metrics, calculated on the validation set.
Train a Gaussian naive Bayes model and a kernel naive Bayes model simultaneously. On the Learn tab, in the Models gallery, click All Naive Bayes. In the Train section, click Train All and select Train Selected.
The app trains one of each naive Bayes model type and highlights the Accuracy (Validation) value of the best model or models. Classification Learner displays a validation confusion matrix for the first model (model 4.1).
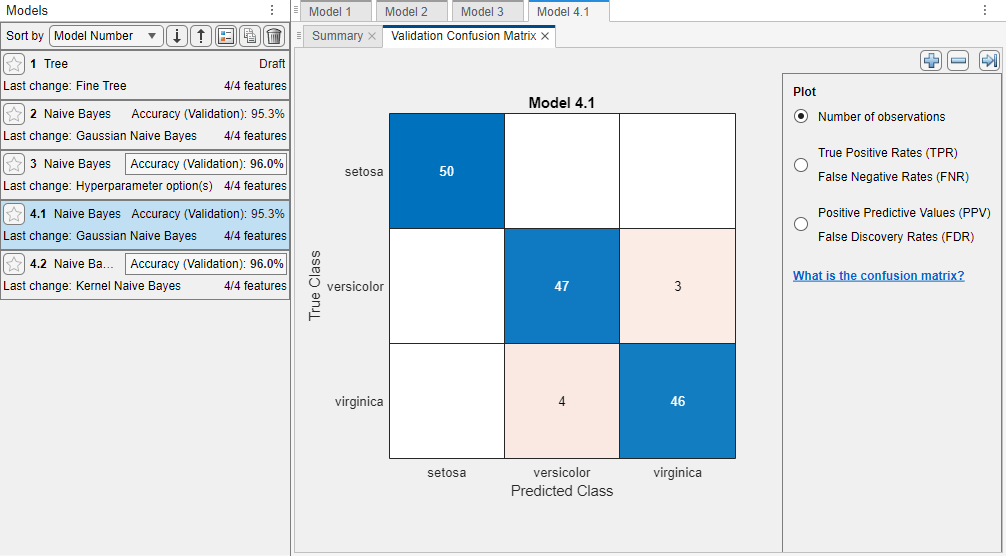
In the Models pane, click a model to view the results. For example, select model 2. Examine the scatter plot for the trained model. On the Learn tab, in the Plots and Results section, click the arrow to open the gallery, and then click Scatter in the Validation Results group. Try plotting different predictors. Misclassified points appear as an X.
Inspect the accuracy of the predictions in each class. On the Learn tab, in the Plots and Results section, click the arrow to open the gallery, and then click Confusion Matrix (Validation) in the Validation Results group. The app displays a matrix of true class and predicted class results.
In the Models pane, click the other trained models and compare their results.
To try to improve the models, include different features during model training. See if you can improve the models by removing features with low predictive power.
On the Learn tab, in the Options section, click Feature Selection.
In the Default Feature Selection tab, you can select different feature ranking algorithms to determine the most important features. After you select a feature ranking algorithm, the app displays a plot of the sorted feature importance scores, where larger scores (including
Infs) indicate greater feature importance. The table shows the ranked features and their scores.In this example, use one-way ANOVA to rank the features. Under Feature Ranking Algorithm, click ANOVA.
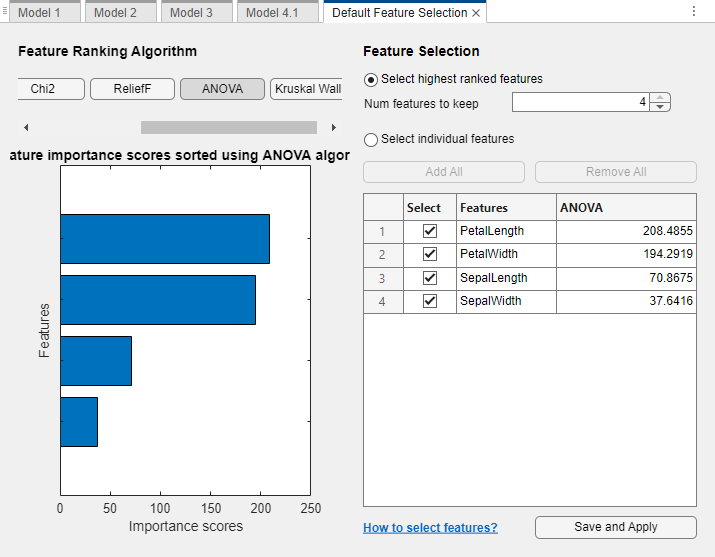
Under Feature Selection, use the default option of selecting the highest ranked features to avoid bias in the validation metrics. Specify to keep 2 of the 4 features for model training. Click Save and Apply. The app applies the feature selection changes to new models created using the Models gallery.
Train new naive Bayes models using the reduced set of features. On the Learn tab, in the Models gallery, click All Naive Bayes. In the Train section, click Train All and select Train Selected.
In this example, the two models trained using a reduced set of features perform better than the models trained using all the predictors. If data collection is expensive or difficult, you might prefer a model that performs satisfactorily without some predictors.
To determine which predictors are included, click a model in the Models pane, and note the check boxes in the expanded Feature Selection section of the model Summary tab. For example, model 5.1 contains only the petal measurements.
Note
If you use a cross-validation scheme and choose to perform feature selection using the Select highest ranked features option, then for each training fold, the app performs feature selection before training a model. Different folds can select different predictors as the highest ranked features. The table on the Default Feature Selection tab shows the list of predictors used by the full model, trained on the training and validation data.
To further investigate features to include or exclude, use the parallel coordinates plot. On the Learn tab, in the Plots and Results section, click the arrow to open the gallery, and then click Parallel Coordinates in the Validation Results group.
In the Models pane, click the model with the highest Accuracy (Validation) value. To try to improve the model further, change its hyperparameters (if possible). First, duplicate the model by right-clicking the model and selecting Duplicate. Then, try changing hyperparameter settings in the model Summary tab. Recall that hyperparameter options are available only for some models. Train the new model by clicking Train All and selecting Train Selected in the Train section.
Export the trained model to the workspace. In the Export section of the Learn tab, click Export Model and select Export Model. In the Export Classification Model dialog box, click OK to accept the default variable name.
Examine the code for training this classifier. On the Learn tab, in the Export section, click Export Model and select Generate Function.
Use the same workflow to evaluate and compare the other classifier types you can train in Classification Learner.
To try all the nonoptimizable classifier model presets available for your data set:
On the Learn tab, in the Models section, click the arrow to open the gallery of models.
In the Get Started group, click All.

In the Train section, click Train All and select Train All.
For information about other classifier types, see Train Classification Models in Classification Learner App.
Related Topics
- Train Classification Models in Classification Learner App
- Select Data for Classification or Open Saved App Session
- Choose Classifier Options
- Naive Bayes Classification
- Feature Selection and Feature Transformation Using Classification Learner App
- Visualize and Assess Classifier Performance in Classification Learner
- Export Classification Model to Predict New Data
