Cluster Data Using Self-Organizing Map (SOM)
This example shows how to train a self-organizing map (SOM) neural network to cluster unlabeled data.
Data clustering is the task of grouping similar observations in a dataset. Typically, clustering is an unsupervised learning task. You do not need a labeled set of training data to train the model. The model does not assign semantic labels to the inputs. The model is trained to output the same group index for similar observations.
A self-organizing map is a type of neural network that performs clustering. It maps high-dimensional data to positions in a lower-dimensional space. The learnable parameters are weight vectors that represent reference points in the space of the training data. During training, the network updates these weight vectors so that similar inputs map to nearby locations in the lower-dimensional space. This process preserves the topological relationships of the original data.
This diagram shows the flow of data through a SOM neural network.
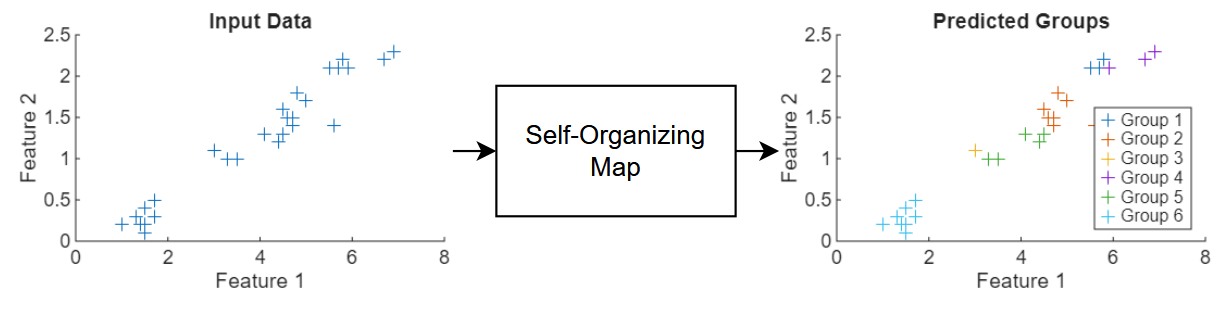
This example trains a SOM neural network that clusters flowers using measurements that correspond to petal and sepal length and width.
Load Training Data
Load the Iris example dataset from iris_dataset. This dataset contains measurements of 150 flowers. The measurements are sepal length, sepal width, petal length, and petal width. Transpose the data so that
load iris_dataset
X = irisInputs';Visualize the data in a plot matrix. The plot in row column is a scatter plot of feature and feature . The histograms on the diagonal show the distribution of the corresponding feature values.
figure
plotmatrix(X)
title("Data Features")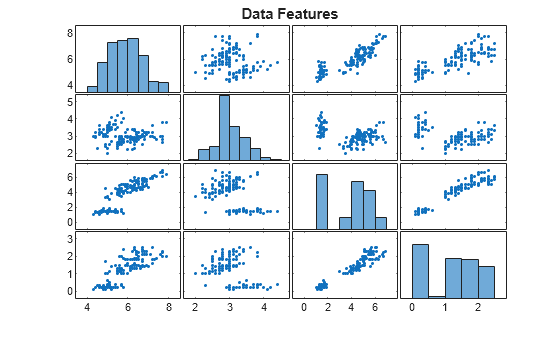
Split the data into training, validation, and test partitions using the trainingPartitions function, which is attached to this example as a supporting file. To access this function, open the example as a live script. Use 80% of the data for training and the remaining 20% for testing.
numObservations = size(X,1); [idxTrain, idxTest] = trainingPartitions(numObservations,[0.8 0.2]); XTrain = X(idxTrain,:); XTest = X(idxTest,:);
Define Neural Network Architecture
Define the neural network architecture for the SOM.
Use a feature input layer with an input size that matches the number of features.
For the SOM operation, use the custom layer
somLayer, attached to this example as a supporting file. to access this layer, open the example as a live script. Use a 3-by-2 grid. To initialize the weights, specify the training data.
gridSize = [3 2];
numFeatures = size(XTrain,2);
layers = [
featureInputLayer(numFeatures)
somLayer(gridSize,XTrain)];Define Model Function
To train the SOM neural network, create a model function that takes the training data as input and returns the normalized neighborhood-augmented activations and the indices of any inactive groups in the map.
The normalized neighborhood-augmented activations represent how strongly the SOM weights respond to the input. They take into account both the closest weight vector and its neighboring nodes. The model uses them to simulate cooperative learning during training, and ensures that similar inputs have similar activations and preserve topological relationships in the map.
function [Y, idxInactive] = model(net,X,iteration,initialNeighborhoodRadius, ... numOrderingIterations,somLayerName) arguments net X iteration initialNeighborhoodRadius numOrderingIterations somLayerName = "som" end % Make predictions. Y = predict(net,X); % Add noise. mask = rand(size(Y)) < 0.9; Y = Y.*mask; % Calculate new radius. r = initialNeighborhoodRadius; N = numOrderingIterations; rNew = 1 + (r-1)*(1 - (iteration - 1)/N); % Determine neighborhood. layer = getLayer(net,somLayerName); distances = layer.NodeDistances; neighborhood = distances <= rNew; % Augment activations and normalize. Y = Y*neighborhood + Y; sumY = sum(Y,1); Y = Y./sumY; % Find inactive nodes. idxInactive = sumY == 0; end
Specify Training Options
Specify the training options. Choosing among the options requires empirical analysis. To explore different training option configurations by running experiments, you can use the Experiment Manager app.
Train the network for 200 iterations.
Train with 100 ordering iterations.
Use an initial neighborhood with a radius size of three.
numIterations = 200; numOrderingIterations = 100; initialNeighborhoodRadius = 3;
Define SOM Update Function
Define a custom update function that updates the SOM weights using the model activations and the indices of the inactive groups.
function net = somupdate(net,X,Y,idxInactive,somLayerName) arguments net X Y idxInactive somLayerName = "som" end % Extract weights. idxSom = net.Learnables.Layer == somLayerName; idxWeights = net.Learnables.Parameter == "Weights"; idx = idxSom && idxWeights; weights = net.Learnables.Value{idx}; % Calculate new weights. newWeights = X'*Y; dWeights = newWeights - weights; dWeights(:,idxInactive) = 0; newWeights = weights + dWeights; % Update learnables. net.Learnables.Value{idx} = dlarray(newWeights); end
Train Neural Network
Train the neural network using a custom training loop.
To make predictions with the neural network, convert the layer array to a dlnetwork object.
net = dlnetwork(layers);
Initialize the training progress monitor.
monitor = trainingProgressMonitor( ... Metrics="QuantizationError", ... Info="Iteration", ... XLabel="Iteration");
Train the neural network. For each iteration:
Calculate the model activations and the indices of the inactive groups using the
modelfunction.Update the learnable parameters using the custom
somupdatefunction.Update the training progress monitor using the
quantizationErrorfunction, listed in the Quantization Error Function section of the example.
iteration = 0; while iteration < numIterations && ~monitor.Stop iteration = iteration + 1; [Y, idxInactive] = model(net,XTrain,iteration, ... initialNeighborhoodRadius,numOrderingIterations); net = somupdate(net,XTrain,Y,idxInactive); layer = getLayer(net,"som"); weights = layer.Weights; qe = quantizationError(weights,XTrain); recordMetrics(monitor,iteration,QuantizationError=qe); updateInfo(monitor, ... Iteration=iteration + " of " + numIterations); monitor.Progress = 100*iteration/numIterations; end
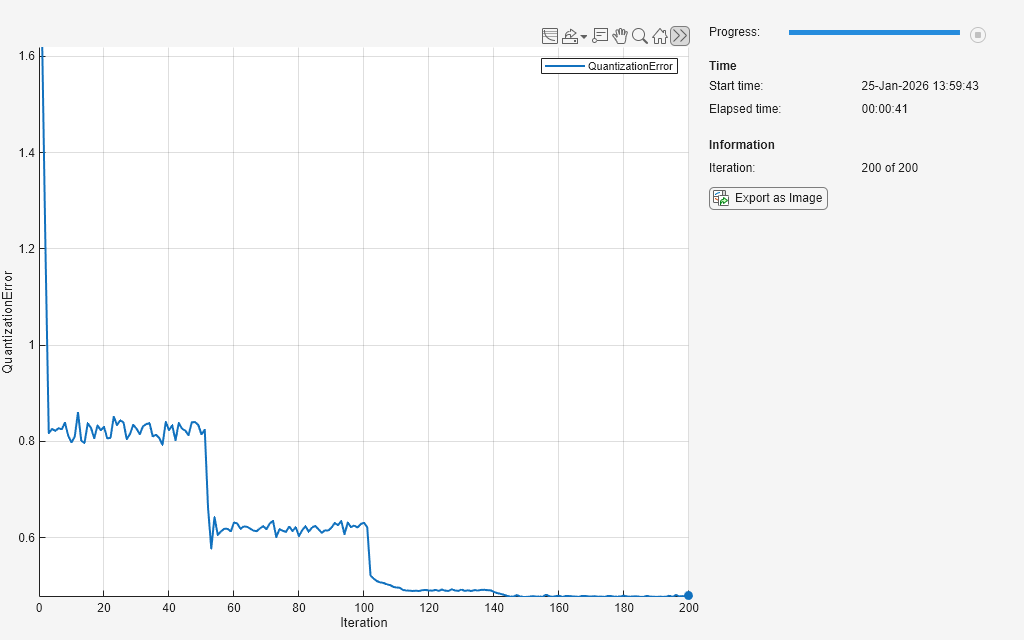
Test Neural Network
Because clustering is an unsupervised learning task, there are no labels to help evaluate the accuracy of the neural network. Instead, you must evaluate the test predictions manually.
Make predictions with the neural network. To convert the network outputs to integer values, use the onehotdecode function and specify the output class type as "double".
numGroups = prod(gridSize);
groupNames = "Group " + string(1:numGroups);
Y = minibatchpredict(net,XTest);
Y = onehotdecode(Y,groupNames,2);Calculate the quantization error.
layer = getLayer(net,"som");
weights = layer.Weights;
qeTest = quantizationError(weights,XTest)qeTest = 0.5543
Visualize the predicted groups against the first two features in a scatter plot.
figure hold on for i = 1:numGroups idx = double(Y) == i; scatter(XTest(idx,1),XTest(idx,2),"+"); end xlabel("Feature 1") ylabel("Feature 2") legend(groupNames,Location="bestoutside")
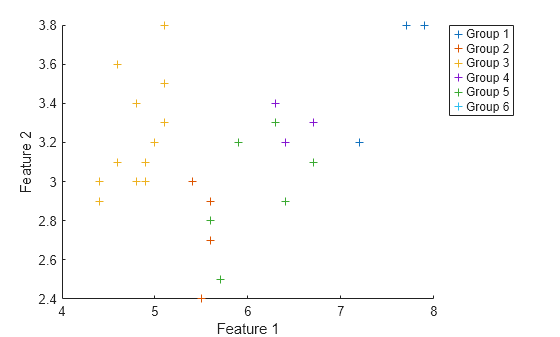
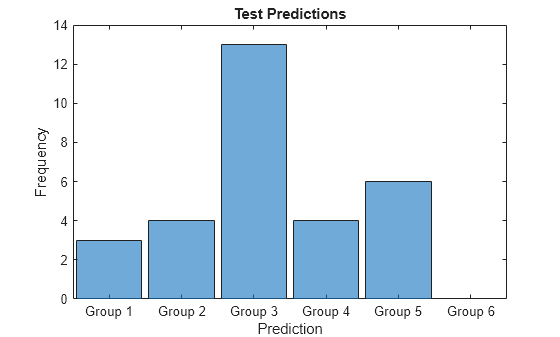
Visualize the distribution of the predicted groups in a histogram.
figure histogram(Y) xlabel("Prediction") ylabel("Frequency") title("Test Predictions")
Visualize the SOM hits in a heatmap. Using the predictions, extract the corresponding group positions from the SOM layer. For each of the nodes in the SOM grid, display the count of the corresponding prediction.
YPositions = layer.Positions(:,Y);
M = histcounts2(YPositions(1,:),YPositions(2,:));
figure
heatmap(M)
title("SOM Hits")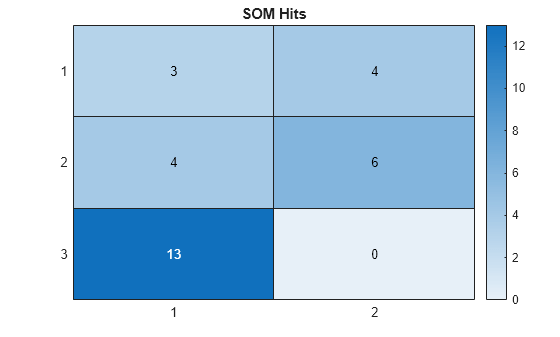
Quantization Error Function
In the context of self-organizing maps, the quantization error measures how well the SOM weights represent the input data.
The quantizationError function takes the SOM weights and the network input data as input and returns the mean of the euclidean distances between each input vector and the closest weight vector. A lower quantization error indicates a closer match between the SOM representation and the input data.
function qe = quantizationError(weights,X) numObservations = size(X,1); minDistances = zeros(numObservations,1); for i = 1:numObservations diffs = weights' - X(i,:); distances = sqrt(sum(diffs.^2,2)); minDistances(i) = min(distances); end qe = mean(minDistances); end
Copyright 2025 The MathWorks, Inc.