pdbdistplot
Visualize intermolecular distances in Protein Data Bank (PDB) file
Description
pdbdistplot( retrieves the structure
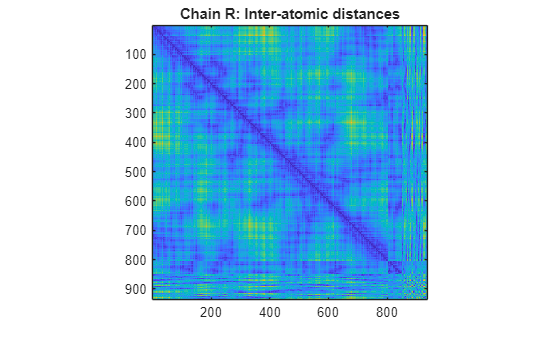
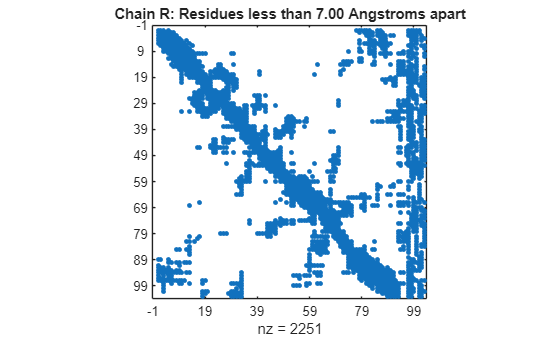
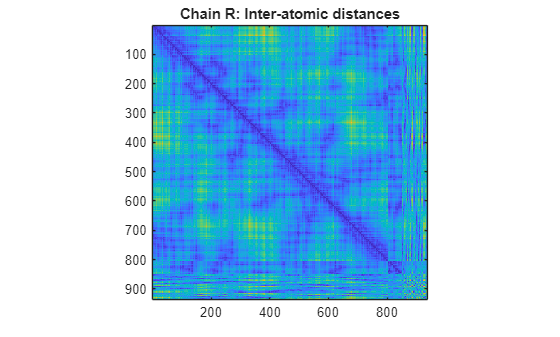
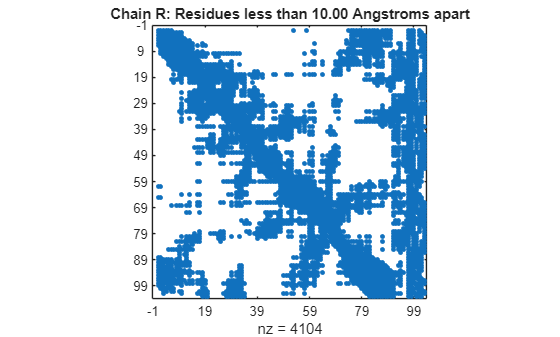
specified by PDBid)PDBid from the PDB database and creates a heat map showing
inter–residue distances and a spy plot showing the residues where the minimum distances
apart are less than 7 angstroms. If multiple chains are present in
PDBid, separate plots are created.
pdbdistplot(___,
specifies additional options using one or more name-value arguments. Use any arguments from
the previous syntaxes.Name=Value)
Examples
Input Arguments
Name-Value Arguments
Version History
Introduced before R2006a
See Also
getpdb | pdbread | proteinplot | ramachandran